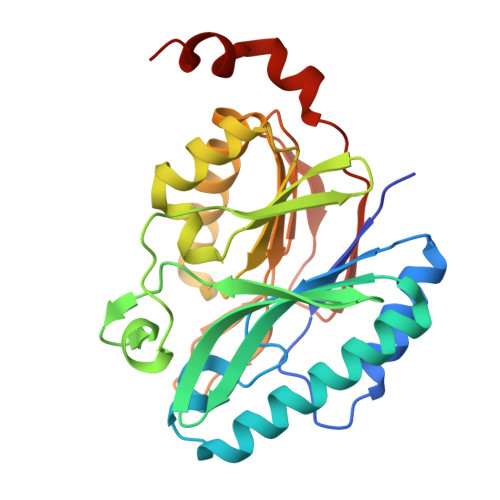
Structural analysis of the unique nitrilase superfamily protein CJ1056C from Campylobacter jejuni.
Ahn, S.Y., Yoon, S.I.(2025) Biochem Biophys Res Commun 785: 152688-152688
- PubMed: 41005285 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2025.152688
- Primary Citation Related Structures:
9WPN, 9WPO - PubMed Abstract:
The nitrilase superfamily comprises enzymes that hydrolyze nonpeptide carbon-nitrogen bonds in nitriles and amides and contribute to diverse cellular processes, such as signaling, metabolism, and detoxification. These enzymes rely on the canonical Cys-Glu-Lys catalytic triad for their nitrilase or amidase activities. We identified a gene product of Campylobacter jejuni (CJ1056C) belonging to the nitrilase superfamily and determined its unique structure by X-ray crystallography in two crystal forms at resolutions of 2.3 Å and 2.9 Å. CJ1056C folds into a four-layered αββα structure with a pocket, as observed in typical nitrilase superfamily proteins, such as Nit2 and aliphatic amidase. However, the catalytic triad of the nitrilase superfamily which is generally located at the base of the pocket, is disrupted in CJ1056C, with the catalytic residues cysteine and lysine replaced by glycine and glutamine, respectively, which are inherently unable to mediate hydrolysis. Furthermore, phylogenetic analysis revealed that CJ1056C and its orthologs from the Campylobacterota phylum form a unique cluster that is evolutionarily separate from catalytically active nitrilase superfamily proteins. Taken together, these findings establish CJ1056C as the first structurally characterized pseudoenzyme of the nitrilase superfamily, presumably lacking conventional nitrilase or amidase activities.
- Division of Biomedical Convergence & Department of Biomedical Science, Kangwon National University, Chuncheon, 24341, Republic of Korea.
Organizational Affiliation: