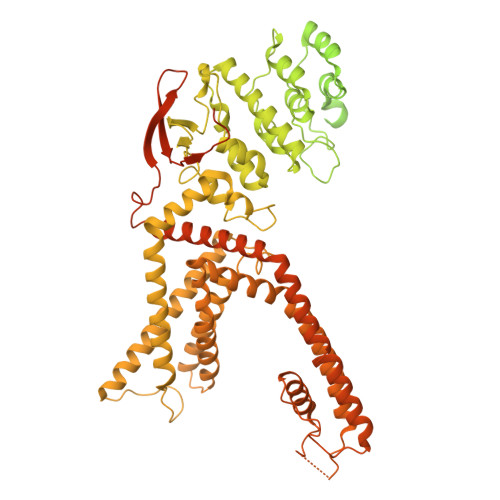
Structures of TRPV1 bound by hyperthermia-inducing analgesics.
Gao, Y.H., Huang, Y.Z., Li, Z.X., Chen, X.Y., Shao, C.Y., Li, H.W., Liu, B., Yang, F., Chen, M.R., Lu, M.L., Zhu, M.X., Yang, F., Xiao, Y.B., Yu, Y.(2026) Cell Rep 45: 116765-116765
- PubMed: 41447532 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2025.116765
- Primary Citation Related Structures:
9W4M, 9W4N, 9W4T - PubMed Abstract:
TRPV1, a member of the transient receptor potential vanilloid subfamily, mediates nociception and thermoregulation. TRPV1-targeting analgesics frequently induce hyperthermia, underscoring the need for structural insights to guide the development of safer compounds. Here, we determined the structures of rat TRPV1 bound to the clinical candidate analgesics AMG517, AMG9810, and SB366791. AMG517 and AMG9810 are deeply situated within the S3-S4 interface of the vanilloid pocket, where they interact with residues from the S3-S6 helices, as well as the S4-S5 linker. These interactions induce local deformations in the TRP-box and lower S6 helix, accompanied by a modest rotation of the S1-S4 bundle, leading to partial dilation of the lower gate. The distinct allosteric changes of AMG517 and AMG9810, compared with the non-hyperthermic ligand SB366791, suggest a structural basis by which TRPV1-targeting analgesics influence thermoregulation and provide insights for designing safer analogs.
- Department of Basic Medicine, Schools of Basic Medicine and Clinical Pharmacy, China Pharmaceutical University, Nanjing 210009, China.
Organizational Affiliation: