Structure of defense against restriction proteins DarA and Hdf in phage P1 reveals a new molecular mechanism during phage assembly, infection and DNA ejection.
Zheng, J., Chen, Y., Chen, S., Zhou, J., Xiao, H., Yang, F., Liu, H.(2026) PLoS Pathog 22: e1013869-e1013869
- PubMed: 41544131 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1013869
- Primary Citation Related Structures:
9UKM, 9UMS, 9VL4, 9VYI, 9VZ0, 9VZK - PubMed Abstract:
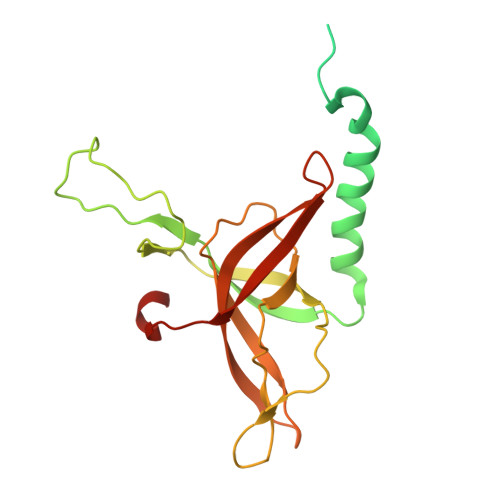
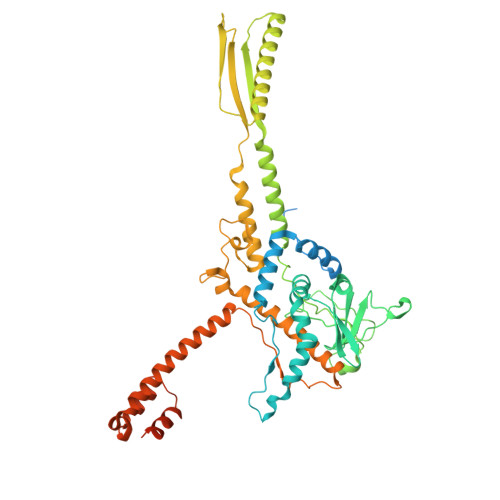
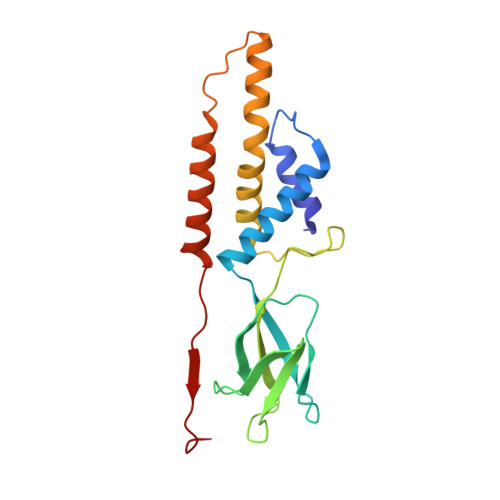
The continuous "arms race" between bacterial antiviral defense systems and phage anti-defense strategies drives evolutionary innovation. Previous study indicated that the defense against restriction (Dar) proteins DarA and Hdf in myophage P1 are associated with the head morphogenesis. However, the structural information for these proteins was lacking, and the mechanisms by which they mediate head morphogenesis and protect phage DNA against bacterial defense systems remained poorly understood. Using cryo-electron microscopy (cryo-EM), we resolved the entire structures of extended P1 and contracted P1 with partial DNA, with the latter lacking the baseplate, as well as the head structure of contracted P1 without DNA. We identified the structural proteins for the P1, including the head, connector complex, and baseplate, which exhibited conserved properties among the majority of myophages with a simple baseplate. Notably, 55 DarA-Hdf pairs are attached to the inner surface of head at each penton-hexon junction in the extended P1 and contracted P1 with partial DNA. The DarA and Hdf together form a complex that is tightly bound to the capsid and interacts with the DNA. However, these pairs are absent in the contracted P1 without DNA. Based on our three states of P1, we hypothesis that these extensive interactions among DarA, Hdf, DNA, and head play crucial roles in mediating capsid assembly, enhancing capsid stability, and protecting phage DNA. Our results provide a structural basis for further exploration of the mechanism by which Dar proteins function during phage assembly, infection and DNA ejection. This molecular mechanism may be conserved among P1-like phages.
- Institute of Interdisciplinary Studies, Key Laboratory for Matter Microstructure and Function of Hunan Province, Key Laboratory of Low-dimensional Quantum Structures and Quantum Control, School of Physics and Electronics, Hunan Normal University, Changsha, China.
Organizational Affiliation: