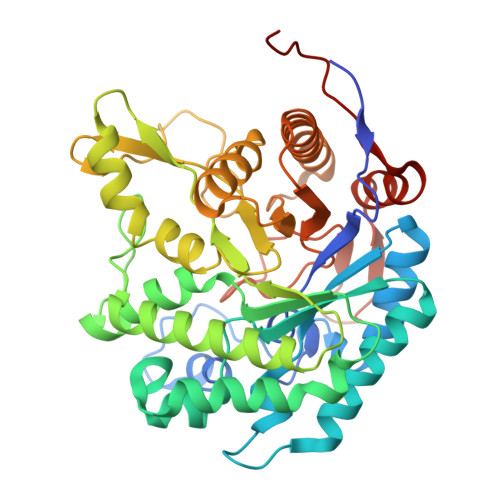
A beta-1,2-glucan-associated glycoside hydrolase family 1 beta-glucosidase from Streptomyces griseus.
Kumakura, H., Motouchi, S., Kobayashi, K., Inoue, M., Kariuda, N., Nakai, H., Nakajima, M.(2025) Protein Sci 34: e70255-e70255
- PubMed: 40838837 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.70255
- Primary Citation Related Structures:
8KAP, 9U98 - PubMed Abstract:
β-Glucosidases, major enzymes that release glucose from various natural compounds, are phylogenetically classified into glycoside hydrolase (GH) families. GH1 is the largest of these families. No β-1,2-glucan-associated GH1 enzyme has been found, even though β-1,2-glucans are natural carbohydrates that are important for interaction between organisms and environmental adaptation. In this study, functional and structural analyses of a GH1 enzyme from Streptomyces griseus (SGR_2426 protein) were performed. SGR_2426 showed the highest hydrolytic activity toward p-nitrophenyl β-glucopyranoside among p-nitrophenyl sugars. This enzyme showed hydrolytic activity toward β-1,2-glucooligosaccharides specifically among β-linked glucooligosaccharides. A structure of the enzyme in complex with sophorose (β-1,2-glucodisaccharide) was obtained as a Michaelis complex. The six-membered ring of the glucose unit at the reducing end of sophorose is positioned in a hydrophobic environment between Trp291 and Met171, while only residue Gln229 forms a hydrogen bond directly. Trp291 and Gln229 are proposed as candidates for the residues important for substrate specificity based on comparison with structurally characterized GH1 homologs. Mutational analysis of Trp291 and Gln229 suggested that Trp291 is important for substrate recognition but not for substrate specificity and that Gln229 is involved in substrate specificity. SGR_2426 is the first identified β-1,2-glucan-associated β-glucosidase in the GH1 family.
- Department of Applied Biological Science, Faculty of Science and Technology, Tokyo University of Science, Noda, Japan.
Organizational Affiliation: