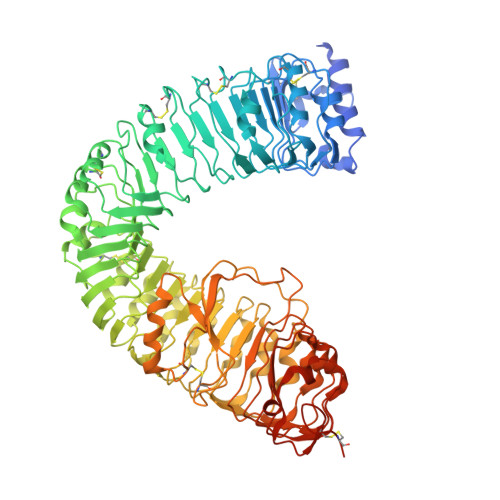
A mechanistic framework for the recognition of chemically diverse brassinosteroids by BRI1-family receptor kinases.
Caregnato, A., Chen, H., Kvasnica, M., Hohmann, U., Oklestkova, J., Ferrer, K., Broger, L., Hothorn, L.A., Strnad, M., Hothorn, M.To be published.