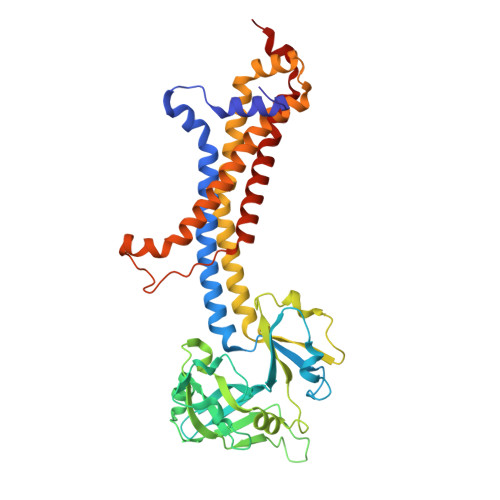
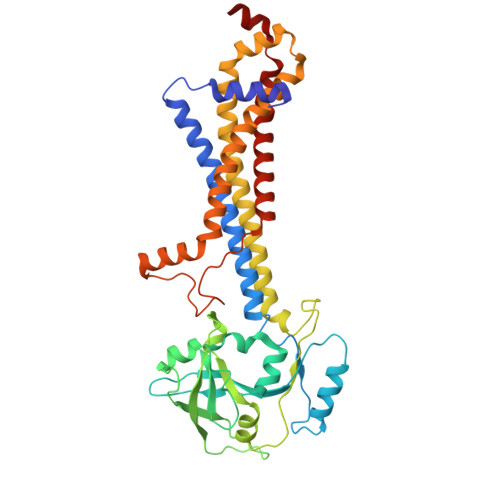
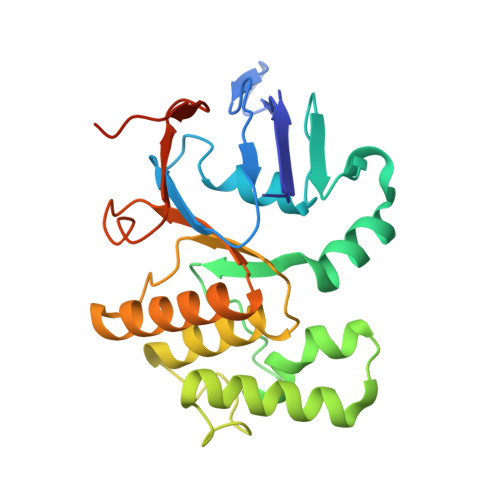
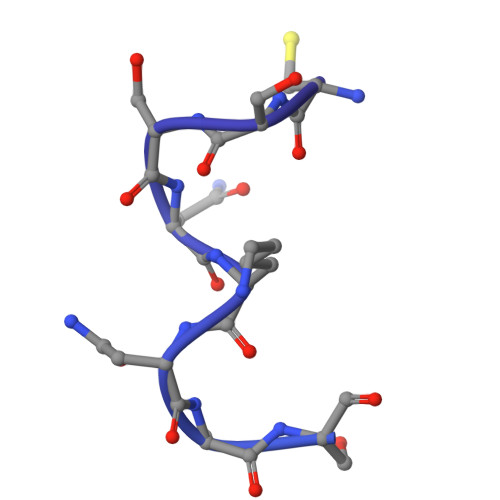
Liganded LolCDE structures reveal a common substrate-LolE interaction guiding bacterial lipoprotein transport.
Szewczyk, P., Greene, N.P., Symmons, M.F., Hardwick, S.W., Koronakis, V.(2026) Proc Natl Acad Sci U S A 123: e2520579123-e2520579123
- PubMed: 41557797 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2520579123
- Primary Citation Related Structures:
9RLC, 9RLD, 9RLE, 9RLF, 9RLG, 9RLH, 9RLI, 9RLJ, 9RLK - PubMed Abstract:
Bacterial lipoproteins are key structural components of the outer membrane in Gram-negative bacteria and vital components of machineries required for its biosynthesis and maintenance. The Lol system, essential for viability, directs transport of lipoproteins from the site of biosynthesis on the inner membrane to the outer membrane and has been the target of extensive efforts to develop novel antimicrobial drugs. In the first stage of this transport process, newly synthesized lipoproteins are released from the inner membrane by the ABC transporter LolCDE and passed to the periplasmic chaperone, LolA. Here, we show cryo-EM structures of LolCDE in complex with three different lipoprotein substrates, Lpp, Pal, and LolB, with the latter two bearing a disordered peptide linker between the acyl chains and the globular domain. Our work reveals that when the mature lipoprotein lacks an unstructured linker, the N-terminal portion of the protein is in an unfolded state for transport. The lipoproteins make a sequence-independent but structurally conserved interaction with a cleft on the surface of the periplasmic domain of LolE that promotes efficient transport. We propose a model of lipoprotein export where this interaction acts as pivot point for the peptide portion of the lipoprotein allowing the acyl chains to rotate 180° from their initial position in LolCDE to their binding site in LolA. Our results demonstrate how LolCDE can extrude lipoproteins of diverse sequence and structure and reveal an important detail of a transport process fundamental to bacterial physiology.
- Department of Pathology, University of Cambridge, Cambridge CB2 1QP, United Kingdom.
Organizational Affiliation: