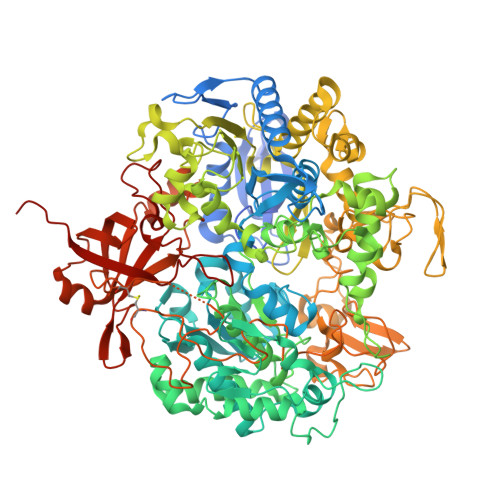
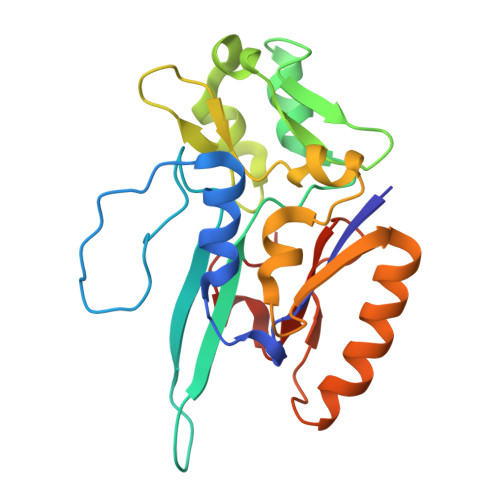
Structural Insights Into CO 2 Transport Pathways in a W-Formate Dehydrogenase: Structural Basis for CO 2 Reduction.
Vilela-Alves, G., Manuel, R.R., Martins, G., Carpentier, P., Raczynska, A., Szaleniec, M., Pereira, I.A.C., Romao, M.J., Mota, C.(2026) Angew Chem Int Ed Engl 65: e26133-e26133
- PubMed: 41787858 Search on PubMed
- DOI: https://doi.org/10.1002/anie.202526133
- Primary Citation Related Structures:
9RJT, 9RJU, 9RJV, 9RJW, 9RJX, 9RJY, 9RJZ, 9RK0, 9RK1 - PubMed Abstract:
Mo/W-dependent formate dehydrogenases (Fdhs) catalyze the reversible reduction of CO 2 to formate and are key biocatalysts with high potential for CO 2 capture/conversion technologies. Although previous studies have suggested the presence of two substrate-access tunnels in Fdhs, experimental evidence for CO 2 -specific pathways has been lacking. Here, we present an integrated study of Nitratidesulfovibrio vulgaris FdhAB combining crystallography, molecular dynamics simulations, mutagenesis, and kinetic assays. NvFdhAB crystals pressurized with Kr, O 2 , and CO 2 were used to map gas diffusion routes and uncovered a substrate-retention site consistently occupied by small molecules in multiple crystal structures. Our results indicate that both substrates mostly use the main tunnel to reach this retention site, but H 2 O and CO 2 can also enter through a novel side branch before following a shared route to the buried W active site. The retention site, located at the junction of both tunnels, plays a synergistic role in enhancing CO 2 reduction by increasing substrate concentration near the catalytic center, thereby improving catalytic efficiency. Notably, variants affecting this site showed a selective effect for CO 2 reduction, with no impact on formate oxidation. These findings provide experimental evidence of a CO 2 -specific pathway and identify structural determinants underpinning efficient CO 2 reduction in this enzyme family.
- UCIBIO, Applied Molecular Biosciences Unit and Associate Laboratory i4HB-Institute for Health and Bioeconomy, Department of Chemistry, NOVA School of Science and Technology, Universidade NOVA de Lisboa, Caparica, Portugal.
Organizational Affiliation: