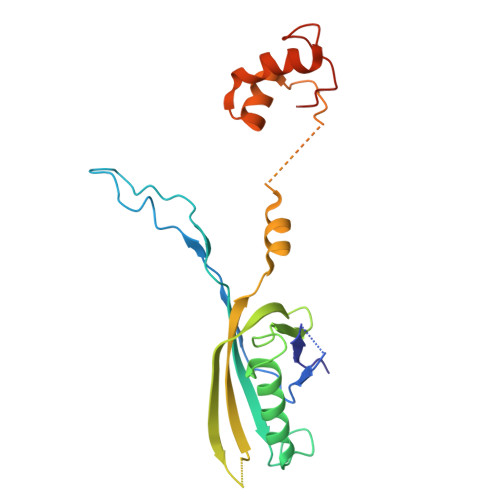
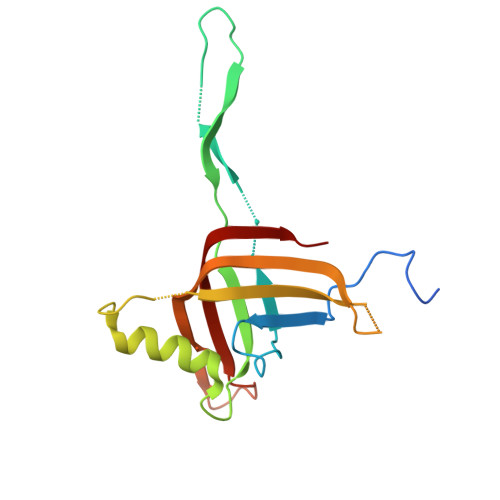
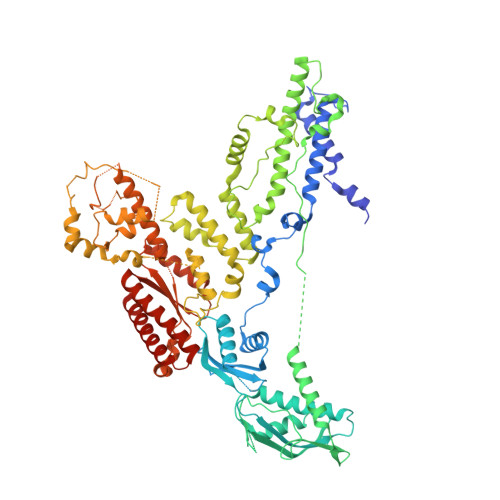
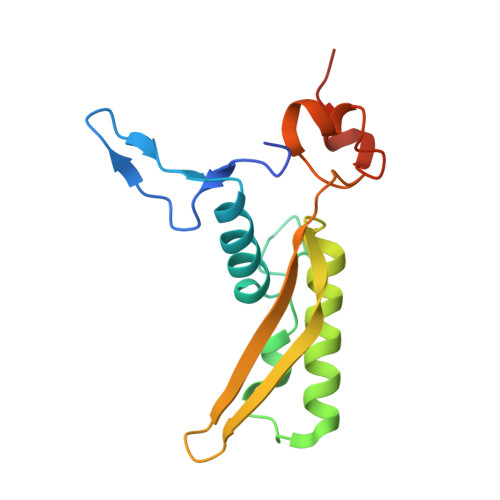
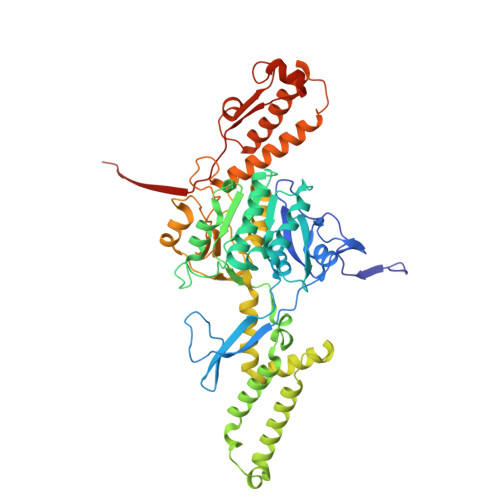
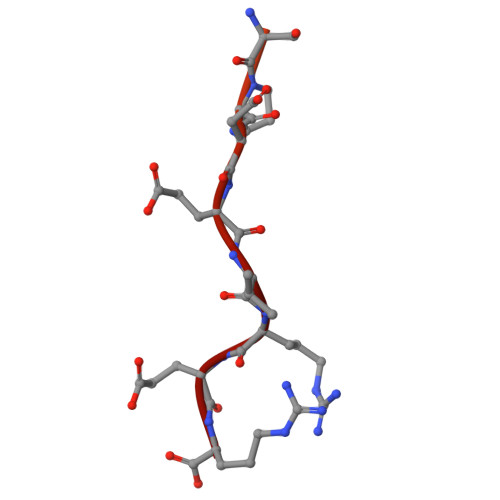
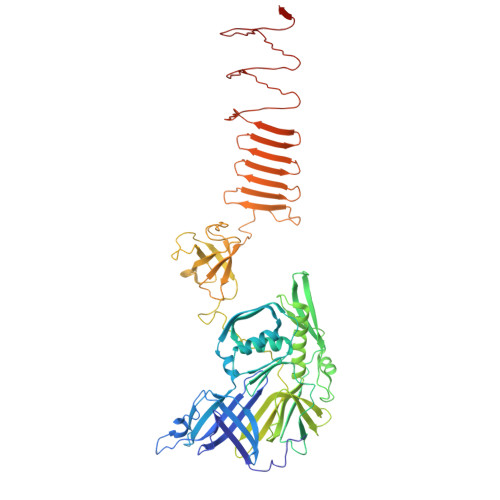
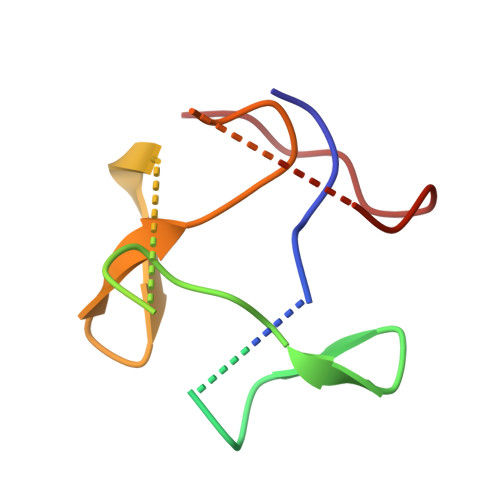
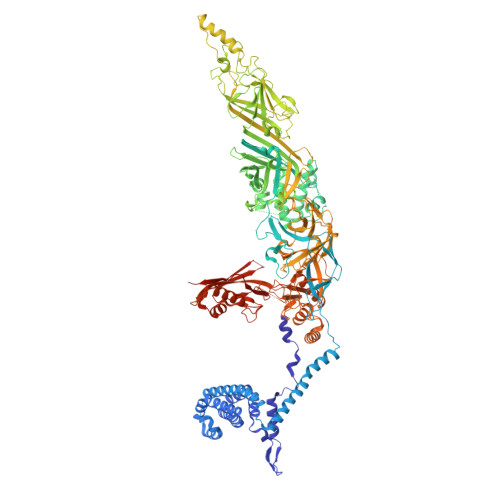
Structure of a contractile injection system in Salmonella enterica subsp. salamae.
Ejaz, R.N., Funke, K., Kielkopf, C.S., Martin, F.J.O., Siborova, M., Hendriks, I.A., Sofos, N.H., Pape, T., Steiner-Rebrova, E.M., Nielsen, M.L., Erhardt, M., Taylor, N.M.I.(2026) Nat Commun
- PubMed: 41991534 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-026-71989-6
- Primary Citation Related Structures:
9R9A, 9R9H, 9R9I, 9R9N, 9RCE - PubMed Abstract:
Extracellular contractile injection systems (eCISs) are phage-derived nanomachines used by bacteria to deliver effectors into target cells. Well-studied examples include the Photorhabdus asymbiotica virulence cassettes and the antifeeding prophage from Serratia entomophila, which have been engineered for heterologous cargo delivery. Recent genomic analyses identified eCIS gene clusters in the opportunistic human pathogen Salmonella enterica subspecies salamae, but their structure, function, and biotechnological potential remain unexplored. Here, we report a high-resolution cryo-electron microscopy structure of the S. enterica eCIS. Our atomic models reveal a distinctive sheath architecture, an expansive cage-like shell around a central spike, and an associated integral membrane protein. We identify a putative effector encoded within the operon exhibiting mild periplasmic toxicity and provide evidence that the S. enterica eCIS deviates from canonical eCISs by interacting with the inner membrane. Guided by these structural features, we uncover, to the best of our knowledge, a previously unannotated cluster of contractile injection systems (CISs). Together, our findings expand the known diversity of CISs' structures and functions, and lay the groundwork for engineering customisable protein delivery platforms.
- Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen, Copenhagen, Denmark.
Organizational Affiliation: