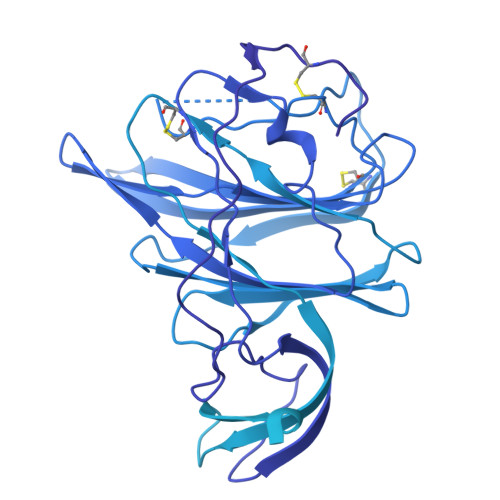
Dipeptidase 1 is a functional receptor for a porcine coronavirus.
Dufloo, J., Fernandez, I., Arbabian, A., Haouz, A., Temperton, N., Gimenez-Lirola, L.G., Rey, F.A., Sanjuan, R.(2025) Nat Microbiol 10: 2981-2996
- PubMed: 41073662 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-025-02111-7
- Primary Citation Related Structures:
9H0B, 9H3J, 9R6O, 9R6P, 9R6Q, 9R6R - PubMed Abstract:
Coronaviruses of the subgenus Embecovirus include several important pathogens, such as the human seasonal coronaviruses HKU1 and OC43, bovine coronavirus and porcine haemagglutinating encephalomyelitis virus (PHEV). While sialic acid is thought to be required for embecovirus entry, protein receptors remain unknown for most of these viruses. Here we show that PHEV does not require sialic acid for entry and instead uses dipeptidase 1 (DPEP1) as a receptor. Cryo-electron microscopy at 3.4-4.4 Å resolution revealed that, unlike other embecoviruses, PHEV displays both open and closed conformations of its spike trimer at steady state. The spike receptor-binding domain (RBD) exhibits extremely high sequence variability across embecoviruses, and we found that DPEP1 usage is specific to PHEV. In contrast, the X-ray structure of the RBD-DPEP1 complex at 2.25 Å showed that the structural elements involved in receptor binding are conserved, highlighting the remarkable versatility of this structural organization in adopting novel receptor specificities.
- Institute for Integrative Systems Biology, CSIC-Universitat de València, Paterna, València, Spain.
Organizational Affiliation: