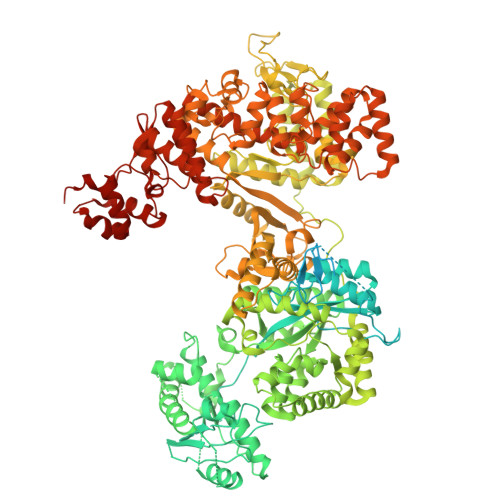
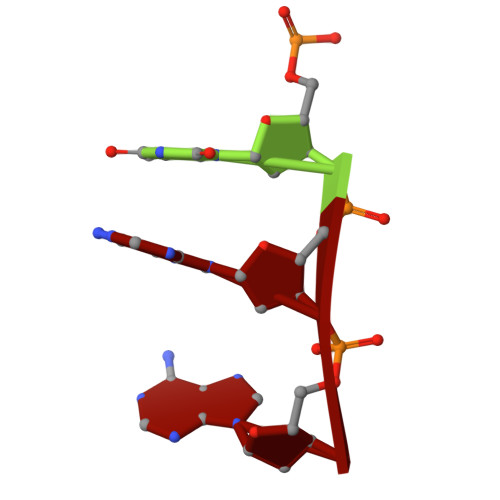
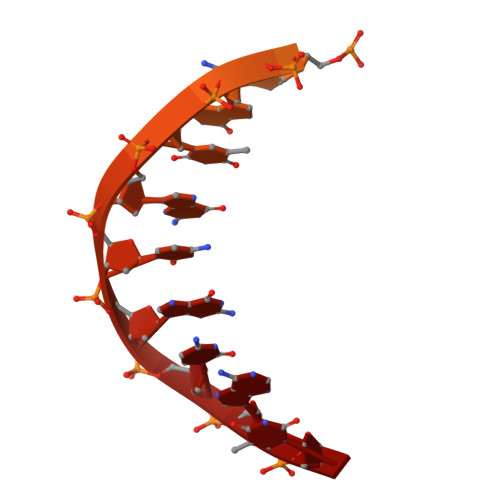
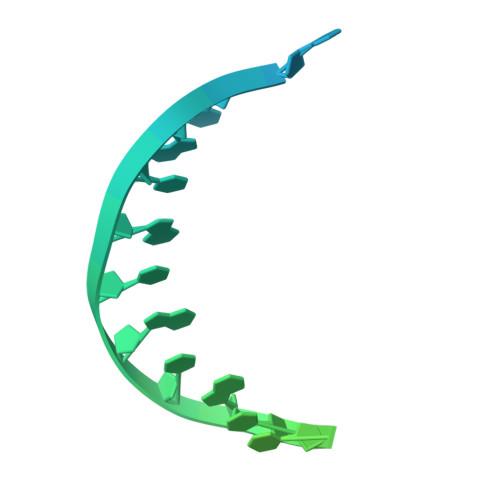
A unique inhibitor conformation selectively targets the DNA polymerase PolC of Gram-positive priority pathogens.
Urem, M., Friggen, A.H., Musch, N., Silverman, M.H., Swain, C.J., Barbachyn, M.R., Mortin, L.I., Yu, X., DeLuccia, R.J., Lamers, M.H., Smits, W.K.(2025) Nat Commun 16: 9784-9784
- PubMed: 41198680 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-65324-8
- Primary Citation Related Structures:
9QPC, 9QRL, 9QRN - PubMed Abstract:
Infections with antimicrobial resistant pathogens are a major threat to human health. Inhibitors of the replicative polymerase PolC are a promising novel class of antimicrobials against Gram-positive pathogens, but the structural basis for their activity remains unknown. The first-in-class PolC-targeting antimicrobial, ibezapolstat, is a guanine analogue in late-stage clinical development for the treatment of Clostridioides difficile infections, and related inhibitors are being developed for systemic treatment of infections with methicillin-resistant Staphylococcus aureus (MRSA) and vancomycin-resistant enterococci (VRE). Here, we present the cryo-electron microscopy structures of Enterococcus faecium PolC bound to DNA and in complex with ibezapolstat or the previously-undescribed inhibitor ACX-801. Both inhibitors form base-pairing interactions with the DNA in the active site, thereby competing with incoming dGTP nucleotides. We identify a crucial susceptibility determinant in PolC that is conserved in other organisms, such as C. difficile. This is explained by an unusual non-planar conformation of the inhibitors that induce a binding pocket in PolC. By combining structural, biochemical, bioinformatic and genetic analyses, this work lays the foundation for the rational development of an innovative class of antimicrobials against Gram-positive priority pathogens.
- Leiden University Center of Infectious Diseases (LUCID), Leiden University Medical Center, Leiden, The Netherlands.
Organizational Affiliation: