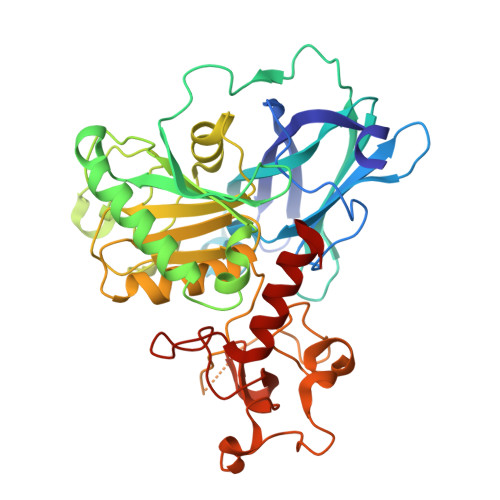
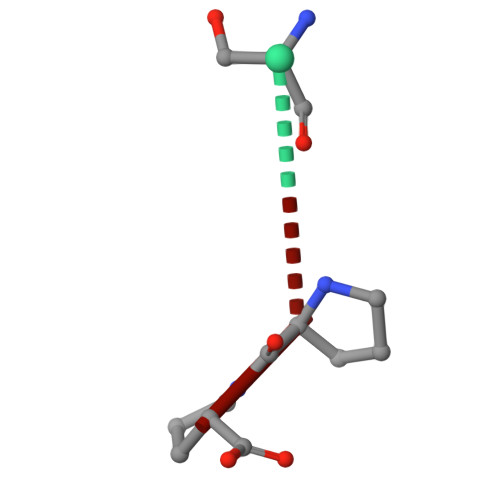
Molecular basis of Fab-dependent IgA antibody recognition by gut-bacterial metallopeptidases.
Marquez-Monino, M.A., Martinez Gascuena, A., Azzam, T., Persson, A., Manzanares-Gomez, A., Aguillo-Urarte, M., Brown, T.T., Montero-Sagarminaga, A., Lood, R., Naegeli, A., Connell, S.R., Sastre, D.E., Sundberg, E.J., Trastoy, B.(2025) EMBO J 44: 4867-4898
- PubMed: 40745064 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44318-025-00518-w
- Primary Citation Related Structures:
9QDH, 9QDI - PubMed Abstract:
Immunoglobulin A (IgA) is essential for mucosal immunity and has been implicated in autoimmune diseases, such as IgA nephropathy. Certain pathogenic and commensal bacteria produce IgA proteases that selectively cleave IgA, potentially aiding bacterial colonization as well as suggesting therapeutic avenues for IgA nephropathy. Here, we investigate the substrate specificities of two enzymes of the M64 metallopeptidase family, the IgA protease ThomasA from Thomasclavelia ramosa and BF3526 from Bacteroides fragilis. Our structural, biochemical, and mutagenesis analyses demonstrate that ThomasA cleaves IgA through exclusive recognition of the Fab region. This mechanism is distinct from that of other antibody-specific peptidases, which typically require engagement of the Fc region. In contrast, X-ray crystal structures of BF3526 in complex with substrate and product peptides, combined with enzymology assays, show that this enzyme targets the N-terminus of pre-digested proteins, but does not act on intact IgA. These findings reveal divergent substrate recognition strategies between M64 family members, while providing new structural insights into their conserved catalytic mechanism.
- Structural Glycoimmunology Laboratory, Biobizkaia Health Research Institute, Barakaldo, Spain.
Organizational Affiliation: