Structure of the Jeilongvirus Receptor Binding Receptor Protein Provides a Molecular Basis for Elongation of the Paramyxoviral C-terminus
Stelfox, A.J., Javorsky, A., Stass, R., Sutton, G., El Omari, K., Bowden, T.A.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Attachment glycoprotein | 435 | J virus | Mutation(s): 0 | 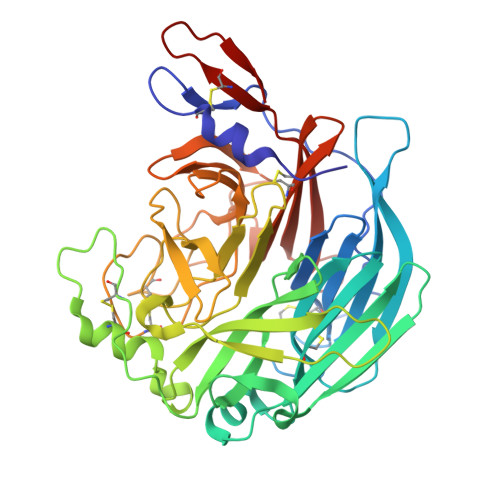 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q49HN3 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 164.24 | α = 90 |
| b = 164.24 | β = 90 |
| c = 112.47 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| xia2 | data reduction |
| xia2 | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Medical Research Council (MRC, United Kingdom) | United Kingdom | MR/L009528/1 |
| Medical Research Council (MRC, United Kingdom) | United Kingdom | MR/S007555/1 |
| Engineering and Physical Sciences Research Council | United Kingdom | EP/K503113/1 |
| Engineering and Physical Sciences Research Council | United Kingdom | EP/L505031/1 |
| Engineering and Physical Sciences Research Council | United Kingdom | EP/M50659X/1 |
| Engineering and Physical Sciences Research Council | United Kingdom | EP/M508111/1 |
| Wellcome Trust | United Kingdom | 203141/Z/16Z |