Single-residue engineering of lambda ( lambda ) antibody light chains reduces conformational flexibility and enhances thermal stability.
Jewel, Y., Young, T., Park, M., Ly, K., Gonzalez, A., Mallett, T.C., Williams, J.C.(2025) Comput Struct Biotechnol J 27: 4730-4739
- PubMed: 41245890 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.csbj.2025.10.045
- Primary Citation Related Structures:
9PZ5, 9PZ6, 9Q0P - PubMed Abstract:
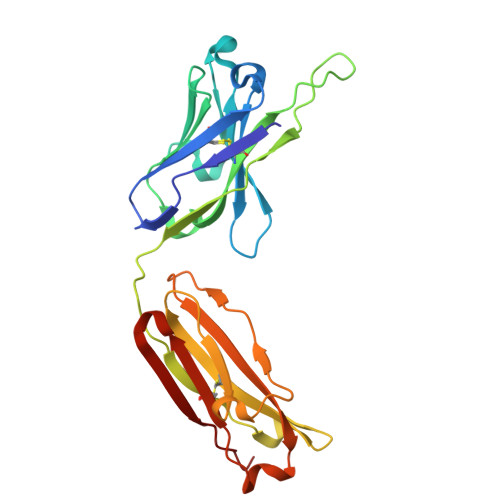
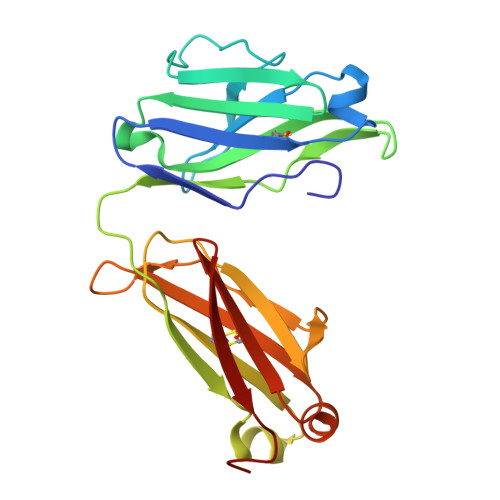
Monoclonal antibodies with lambda (λ) light chains are less commonly used in therapeutics due to their lower biophysical stability compared to kappa (κ) variants. Here, we identify a conserved glycine residue (Gly111) in the λ light chain hinge as a driver of large-scale Fab elbow-angle transitions. Using microsecond-scale molecular dynamics simulations of the EBV-neutralizing Fab AMMO1, we show that substituting Gly111 with threonine (G111T) increases the free energy barrier between conformational states, effectively arresting these transitions. Structural and biophysical analyses-including crystallography, differential scanning fluorimetry, and surface plasmon resonance-confirm that the mutation maintains Fab architecture and antigen binding while increasing thermal stability by up to 2.5 °C. The same mutation applied to a second λ-Fab yielded similar stabilization, and simulations of three clinical λ-Fabs revealed consistent reductions in elbow-angle flexibility. These results demonstrate a generalizable, single-residue engineering strategy to enhance the stability of λ-based Fabs without compromising function, with direct implications for therapeutic antibody development and manufacturability.
- Department of Cancer Biology and Molecular Medicine, Beckman Research Institute, City of Hope National Medical Center, Duarte, CA, United States.
Organizational Affiliation: