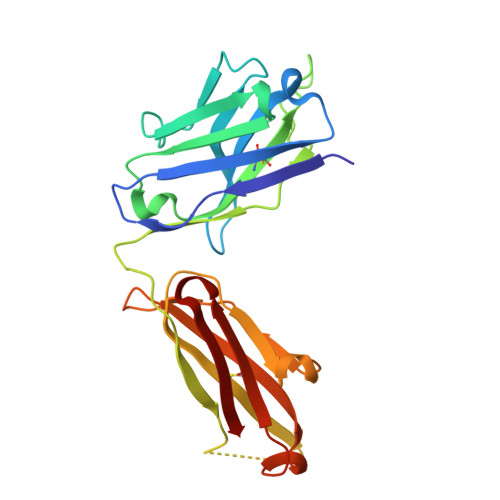
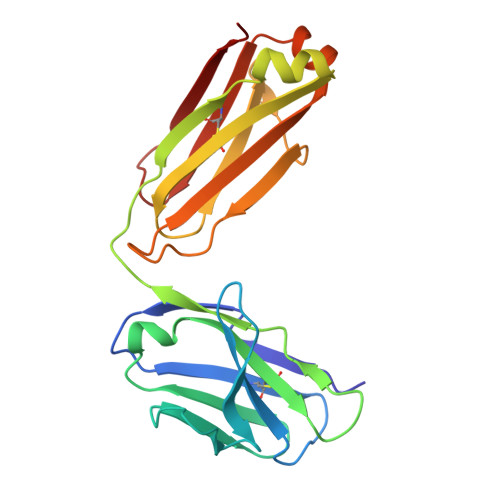
Building a potent TREM2 agonistic, biparatopic, common light chain antibody.
da Silva Almeida, A., Geddie, M.L., Bhate, A., Quan, C., Arndt, J.W., Jiao, Y., Santoro, J.C., Noiman, L., Vijayakumar, R., Sanchez-Salazar, J., Datta, A., Antognetti, G., Hartana, C.A., Wang, X.F., Smith, B.A., Bartlett, D., Duncan, D., Liu, C.C., Otero Gutierrez, K., Cameron, T.O., Koirala, S., Cooke, H.A.(2025) MAbs 17: 2546554-2546554
- PubMed: 40803893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/19420862.2025.2546554
- Primary Citation Related Structures:
9PWN, 9PX5 - PubMed Abstract:
Triggering Receptor Expressed on Myeloid cells 2 (TREM2) plays an important role in microglial function and has been genetically linked to Alzheimer's disease. Activation of TREM2 signaling may contribute to protection against neurotoxic effects of amyloid. Numerous TREM2 activating antibodies have been shown to modulate downstream microglial functions to different degrees, with mixed results in preclinical models and in the clinic. We sought to generate an effectorless agonistic antibody that acted solely through TREM2 engagement with sufficient potency to activate TREM2 in the brain. Our approach focused on building a multivalent biparatopic TREM2 antibody that could mimic the higher order clustering induced by native polyanionic ligands of TREM2. We describe our screening strategy and findings that led to the discovery of a potential therapeutic molecule composed of antibodies selected for optimal affinity, binding epitopes, and geometry. The most productive antibody pair was selected from a common light chain yeast-display library, which required multiple rounds of affinity maturation. Lead antibody candidates were converted into asymmetric tetravalent bispecifics via controlled Fab-arm exchange and subsequently screened in signaling assays. The most productive antibody pair was reengineered into a symmetric tetravalent format, increasing potency and simplifying development. This molecule exhibited higher efficacy and potency in signaling assays than other antibody formats tested and elicited TREM2-mediated chemokine responses in vivo. Our results demonstrate a biparatopic strategy for producing a high potency TREM2 agonistic antibody with low effector function that can modulate TREM2 signaling in vitro and brain pharmacodynamic responses in vivo.
- Biologics Drug Discovery, Biogen, Cambridge, MA, USA.
Organizational Affiliation: