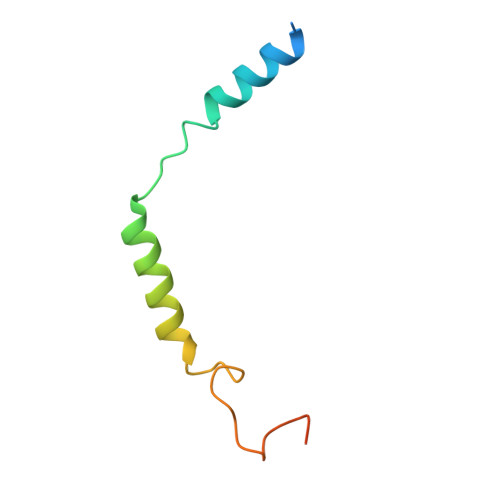
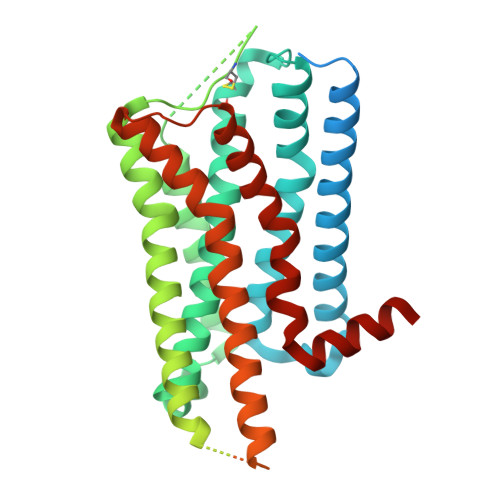
Toward a Random Background for Ligand Optimization
Xu, X., Mailhot, O., Correy, G., Huang, X.P., Braz, J., Shi, D., Srinivasan, K., Holota, Y., Kuziv, Y., Tsoutsouvas, C., Levinzon, N., Doruk, Y.U., Rachman, M., Diolaiti, M., Stevens, M., Liu, F., Hubner, H., Wang, J., Wu, Y., Ashworth, A., Makriyannis, A., Zhang, Y., Moroz, Y., Gmeiner, P., Abel, R., Manglik, A., Basbaum, A., Roth, B., Fraser, J., Shoichet, B.To be published.