Alpha1/Beta Heteromeric Glycine receptor in the presence of 0.200 mM strychnine and 500 nM ivermectin
Gibbs, E., Feddersen, B., Kindig, K.J., Seiferth, D., Biggin, P.C., Chakrapani, S.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Glycine receptor, alpha 1 | A, B [auth D], C, D [auth B] | 458 | Danio rerio | Mutation(s): 0 Gene Names: glra1 | 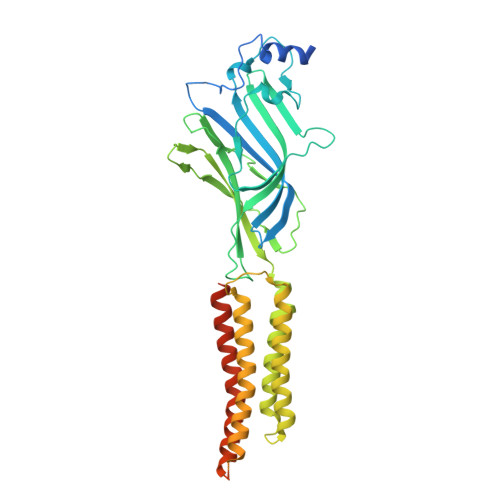 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | B3DHZ5 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Glycine receptor beta subunit 2 | 591 | Danio rerio | Mutation(s): 0 Gene Names: glrbb, glrb2 | 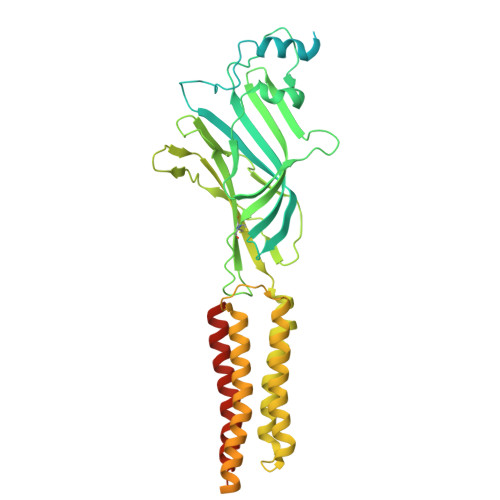 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q6DC22 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SY9 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | F [auth A], I [auth C], K [auth B], M [auth E], N [auth E] | STRYCHNINE C21 H22 N2 O2 QMGVPVSNSZLJIA-FVWCLLPLSA-N |  | ||
| NAG Download:Ideal Coordinates CCD File | G [auth A], H [auth D], J [auth C], L [auth B], O [auth E] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | 1.19.2_4158 |
| RECONSTRUCTION | RELION |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | -- |