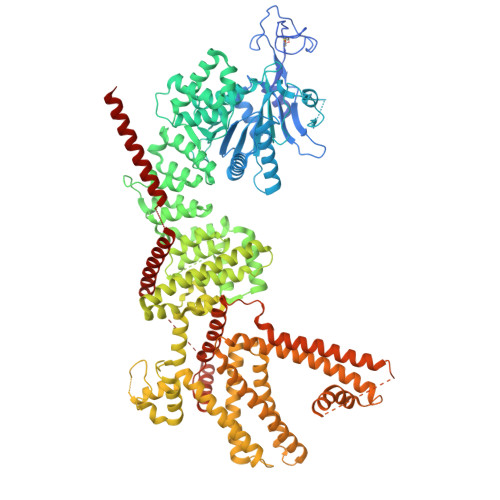
Structural energetics of cold sensitivity.
Choi, K.Y., Lin, X., Cheng, Y., Julius, D.(2026) Nature 653: 962-970
- PubMed: 41882351 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-026-10276-2
- Primary Citation Related Structures:
9P7S, 9P8Y, 9P90, 9P91, 9PAR, 9PB5, 9PB6, 9ZCN, 9ZCO, 9ZCP, 9ZCQ, 9ZCR, 9ZCU, 9ZCV, 9ZEZ, 9ZF0 - PubMed Abstract:
Thermosensitive transient receptor potential (TRP) ion channels enable somatosensory nerve fibres to detect changes in our thermal environment over a wide physiologic range 1-3 . In mammals, the menthol receptor, TRPM8, is activated by temperatures below approximately 26 °C and is essential for the perception of cold or chemical cooling agents 4-6 . A fascinating, yet still unachieved goal is to elucidate mechanisms, both structural and thermodynamic, whereby TRPM8 or other thermosensitive channels are gated by changes in ambient temperature. Recent studies using cryogenic electron microscopy have attempted to address this challenging question but are limited by difficulties in visualizing temperature-evoked conformational sub-states or assessing the energetic landscape governing gating transitions 7,8 . Here we close this gap by combining cryogenic electron microscopy with hydrogen-deuterium exchange mass spectrometry to elucidate a mechanism for cold-evoked activation of TRPM8. First, we visualize TRPM8 channels in cellular membranes, where bona fide menthol- and cold-evoked open states are captured. We also identify a new 'semi-swapped' architecture in which interdigitation of channel sub-units is rearranged substantially following repositioning of the S6 transmembrane helix and elements of the pore region. We then use hydrogen-deuterium exchange mass spectrometry to pinpoint the pore and TRP helices as the regions exhibiting the greatest stimulus-evoked energetic changes that drive channel gating. Specifically, cold-evoked stabilization of the outer pore region repositions the pore lining S6 transmembrane helix while enabling binding of a regulatory lipid to stabilize the open channel. Structural mechanisms associated with activation are validated by comparison of human TRPM8 with the menthol-sensitive but relatively cold-insensitive avian orthologue. We propose a free energy landscape and conformational pathway whereby cold or cooling agents activate this thermosensory receptor.
- Department of Biochemistry and Biophysics, University of California San Franscisco, San Francisco, CA, USA.
Organizational Affiliation: