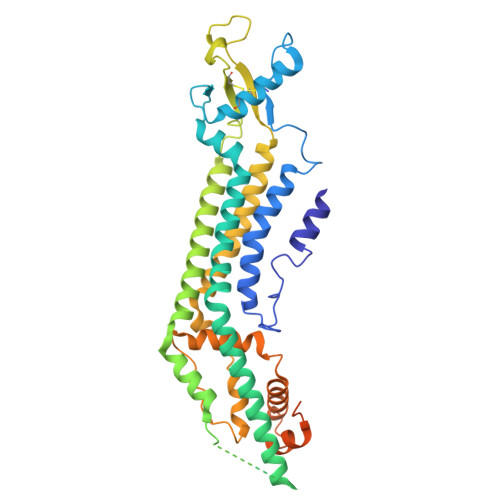
Structural basis of PANX1 permeation and positive modulation by mefloquine.
Li, Y., Ruan, Z., Lee, J., Orozco, I.J., Zhou, E., Du, J., Lu, W.(2025) Nat Commun 16: 11057-11057
- PubMed: 41381453 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-66028-9
- Primary Citation Related Structures:
9OQG, 9OQH, 9OQI, 9OQJ, 9OQK, 9OQL, 9OQM, 9OQN, 9OQO, 9OQP, 9OQQ - PubMed Abstract:
Purinergic signaling relies on ATP release through exocytosis and large-pore channels. Large-pore channels permeate both small anions like chloride and large signaling molecules like ATP, but how this broad cargo selectivity is structurally controlled remains elusive. Here we investigate PANX1, a prototypical large-pore channel, and uncover structural plasticity at the extracellular entrance formed by seven tryptophan (W74) residues. The W74 sidechains are flexible, sampling conformations that range from a constricted state permissive only to chloride to a dilated state compatible with ATP. These states are coupled to variable cation-π interactions between W74 and arginine 75 (R75), suggesting a mechanism for dynamic tuning of pore architecture and selective cargo permeation. We also identify mefloquine as a positive modulator of PANX1 that binds near the side tunnel to control ion flow through this pathway. Together, these findings define the structural principles underlying PANX1 permeation and modulation.
- Department of Molecular Biosciences, Northwestern University, Evanston, IL, USA.
Organizational Affiliation: