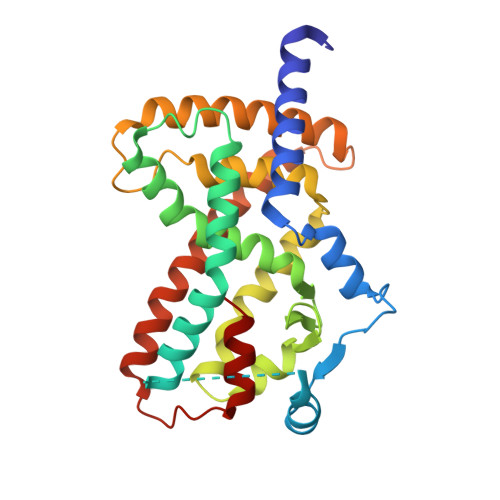
Structural Basis of PPAR gamma-Mediated Transcriptional Repression by the Covalent Inverse Agonist FX-909.
Laughlin, Z.T., Arifova, L., Munoz-Tello, P., Yu, X., Giridhar, M.N.K., Dong, J., Harp, J.M., Zhu, D., Kamenecka, T.M., Kojetin, D.J.(2025) J Med Chem 68: 17587-17597
- PubMed: 40797371 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.5c01252
- Primary Citation Related Structures:
9O9N - PubMed Abstract:
Hyperactivation of peroxisome proliferator-activated receptor γ-mediated transcription promotes tumor growth in urothelial (bladder) cancer, which can be inhibited by compounds that repress PPARγ activity. FX-909 is a covalent PPARγ inverse agonist in phase 1 clinical trials for advanced solid malignancies, including muscle-invasive bladder cancer. Here, we compared the mechanism of action of FX-909 to other covalent inverse agonists including T0070907, reported more than 20 years ago and misclassified as an antagonist, and two reported improved covalent inverse agonist analogs, SR33068 and BAY-4931. Functional profiling and NMR studies reveal that FX-909 displays improved corepressor-selective inverse agonism and better stabilizes a transcriptionally repressive PPARγ LBD conformation compared to T0070907. The crystal structure of PPARγ LBD cobound to FX-909 and the NCoR1 corepressor peptide reveals a repressive conformation shared by other covalent inverse agonists. These findings build on recent studies highlighting the pharmacological significance and clinical relevance of transcriptionally repressive PPARγ inverse agonists.
- Department of Biochemistry, Vanderbilt University, Nashville, Tennessee 37232, United States.
Organizational Affiliation: