The molecular basis of T cell receptor recognition of citrullinated tenascin-C presented by HLA-DR4.
Dao, H.T., Loh, T.J., Sharma, R.K., Klareskog, L., Malmstrom, V., Reid, H.H., Rossjohn, J., Lim, J.J.(2025) J Biological Chem 301: 110326-110326
- PubMed: 40466907 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2025.110326
- Primary Citation Related Structures:
9NIG, 9NIH, 9NII - PubMed Abstract:
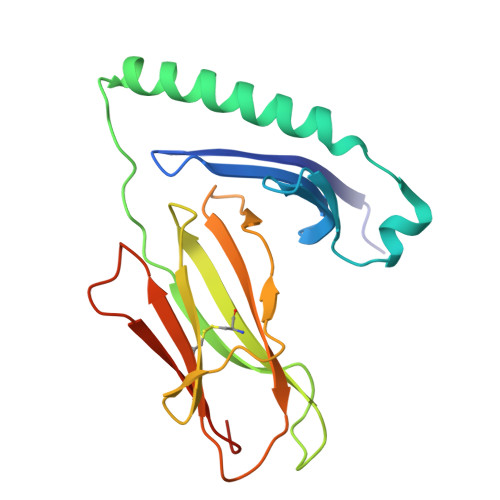
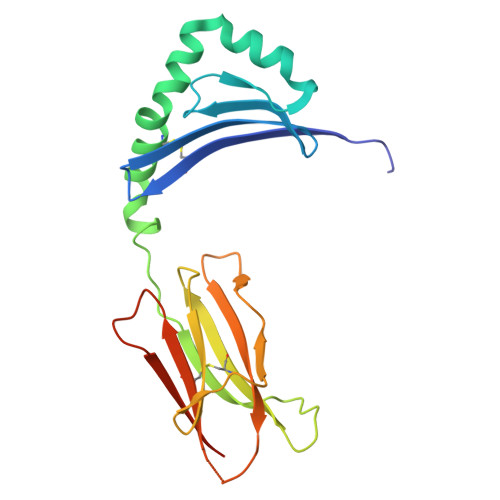
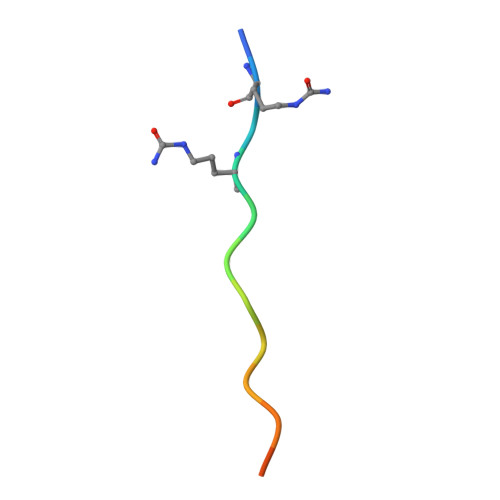
CD4 + T cell autoreactivity against citrullinated (cit) self-epitopes presented by HLA-DRB1 is associated with rheumatoid arthritis (RA) pathogenesis. We understand the molecular bases of T cell receptor (TCR) recognition of cit-fibrinogen, cit-vimentin, and cit-α-enolase epitopes, and the role of citrulline in shaping TCR repertoire usage. Nevertheless, how TCRs recognize other cit-epitopes, including tenascin-C (TNC) and how alternative citrullination positions may modulate the T cell recognition remains unclear. Here, we examined TNC 1014,1016cit peptide, which contains citrullination at position P-1 and P2, to study the underlying TCR-HLA-DRB1 ∗ 04:01-TNC 1014,1016cit molecular interactions. Crystal structure of HLA-DRB1 ∗ 04:01 TNC1014,1016cit at 2.4 Å resolution revealed a conserved peptide binding register to the established HLA-DRB1 ∗ 04:01-peptide structures, where both citrullines protruded upward. Next, we determined the crystal structure of a RA patient-derived TRAV35 + /TRBV10-2 + (PB) TCR in complex with HLA-DRB1 ∗ 04:01 TNC1014,1016cit at 3.2 Å resolution. The CDR3α loop ( 109 VGNTN 113 ) of PB TCR formed a secondary helical conformation at the N-terminus of the peptide binding cleft, allowing extensive interactions between the P-1 and P2 citrullines of TNC 1014,1016cit peptide. Surface plasmon resonance, tetramer staining, and CD69 activation assays revealed that the PB TCR did not cross-react to other RA autoantigens, and the P-1-Cit, P2-Cit, and P5-Tyr of TNC 1014,1016cit are the key determinants underlying the strict specificity of the PB TCR. Collectively, we provide molecular insight into citrullination in modulating TCR recognition.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Australia.
Organizational Affiliation: