Mechanistic basis for a novel dual-function Gag-Pol dimerizer potentiating CARD8 inflammasome activation and clearance of HIV-infected cells.
Hollander, K., Devarkar, S.C., Tang, S., Tiwari, R., Ma, S., Lee, W.G., Denn, E., Wang, Q., Spasov, K.A., Robbins, J.A., Frey, K.M., Jorgensen, W.L., Xiong, Y., Shan, L., Anderson, K.S.(2025) NPJ Drug Discov 2: 22-22
- PubMed: 40904837 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44386-025-00025-2
- Primary Citation Related Structures:
9MXB, 9MXQ, 9MXR, 9MXS, 9MXT - PubMed Abstract:
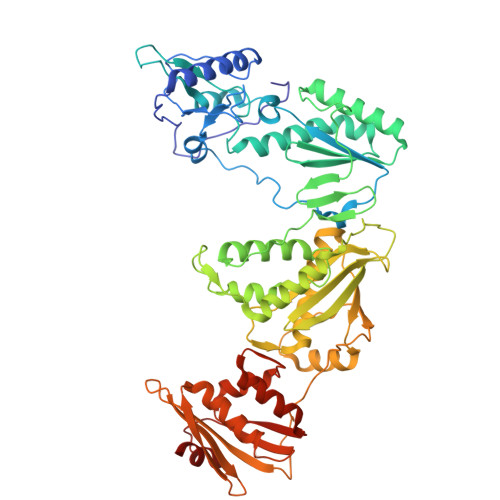
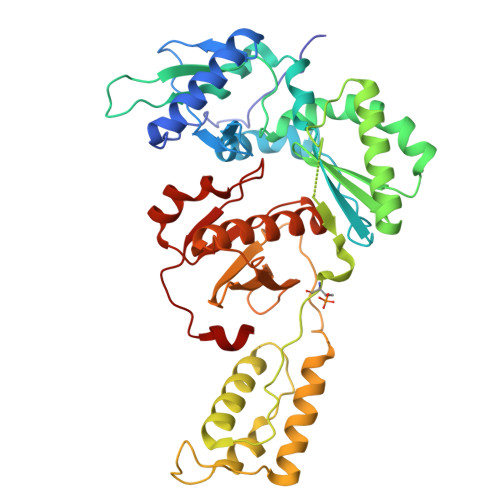
A strategy to functionally cure AIDS by eliminating latent HIV-1 reservoirs involves non-nucleoside reverse transcriptase inhibitors (NNRTIs) that promote pyroptosis of HIV-1 infected cells. These NNRTIs stimulate dimerization of the Gag-Pol polyprotein, resulting in premature HIV-1 protease (PR) dimerization and cleavage of intracellular CARD8. A unique cell-based high-throughput screen was developed to identify potent compounds activating the CARD8 inflammasome through Gag-Pol dimerization. Our in-house library of NNRTIs was evaluated, including a series of catechol diethers, which are potent, nontoxic antivirals. JLJ648 was identified as a promising dual-function antiviral and Gag-Pol dimerizer. Cryo-EM studies of HIV reverse transcriptase p66 bound to JLJ648 revealed populations of homodimers and, surprisingly, a homotetramer. This novel homotetramer structure resembling an 'infinity knot' revealed two JLJ648-bound homodimers forming an extensive interface and nucleated around a dimer of JLJ648 molecules. Structure-guided mutagenesis studies indicate that Gag-Pol homotetramerization may play a critical role in facilitating PR self-cleavage and triggering pyroptosis.
- Department of Molecular Biophysics and Biochemistry, Yale University School of Medicine, New Haven, CT USA.
Organizational Affiliation: