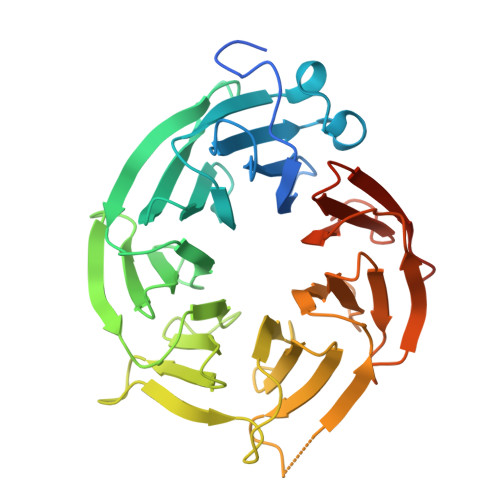
From DNA-Encoded Library Screening to AM-9747 : An MTA-Cooperative PRMT5 Inhibitor with Potent Oral In Vivo Efficacy.
Sarvary, I., Vestergaard, M., Moretti, L., Andersson, J., Peiro Cadahia, J., Cowland, S., Flagstad, T., Franch, T., Gouliaev, A., Husemoen, G., Jacso, T., Kronborg, T., Kuropatnicka, A., Nadali, A., Madsen, M., Nielsen, S.R., Pii, D., Ryborg, S.R., Soede, C., Allen, J.R., Bourbeau, M., Li, K., Liu, Q., Lo, M.C., Madoux, F., Mardirossian, N., Moriguchi, J., Ngo, R., Peng, C.C., Pettus, L., Tamayo, N., Wang, P., Kapoor, R., Belmontes, B., Caenepeel, S., Hughes, P., Liu, S., Slemmons, K.K., Yang, Y., Xie, F., Ghimire-Rijal, S., Mukund, S., Glad, S.(2025) J Med Chem 68: 6534-6557
- PubMed: 40102181 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.4c03101
- Primary Citation Related Structures:
9MRE - PubMed Abstract:
Inhibition of the methyltransferase enzyme PRMT5 by MTA accumulation is a vulnerability of MTAP-deleted cancers. Herein, we report the discovery and optimization of a quinolin-2-amine DEL hit that cooperatively binds PRMT5:MEP50 and MTA to generate a catalytically inhibited ternary complex. X-ray crystallography confirms quinolin-2-amine binding of PRMT5 glutamate-444, while simultaneously exhibiting a hydrophobic interaction with MTA. Lead optimization produced AM-9747 , which selectively inhibits PRMT5-directed symmetric dimethylation of arginine residues of proteins, leading to a potent reduction of cell viability in MTAP-del cells compared to MTAP-WT cells. Once-daily oral dosing of AM-9747 in mouse xenografts is well tolerated, displaying a robust and dose-dependent inhibition of symmetric dimethylation of arginine in MTAP-del tumor-xenografts and significant concomitant tumor growth inhibition without any significant effect on MTAP-WT tumor xenografts.
- Amgen Research, Amgen Inc, Ro̷nnegade 8, DK-2100 Copenhagen, Denmark.
Organizational Affiliation: