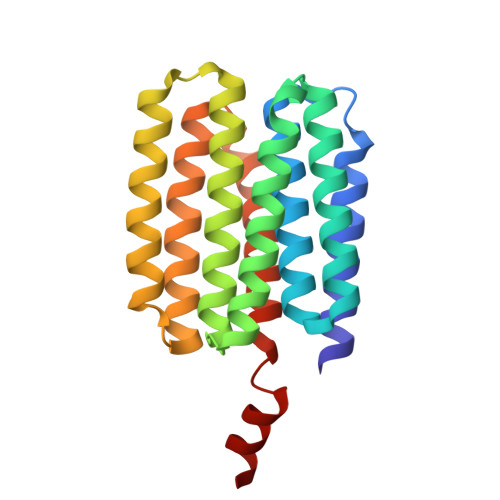
Bioinformatics classification of the MgtE Mg2+ channel and de novo protein design for the stabilization of its novel subclass
Zhao, Z., Omae, K., Iwasaki, W., Zhang, Z., Pan, F., Lee, E., Ito, K., Hattori, M.(2026) Acta Biochim Biophys Sin (Shanghai)