Cryo-EM structure of the ICT01-BTN2A1/BTN3A1/BTN3A2 complex at 4 Angtroms resolution (2 fabs)
Xin, W.Z., Huang, B.D., Zhou, Q.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Butyrophilin subfamily 2 member A1 | 534 | Homo sapiens | Mutation(s): 0 Gene Names: BTN2A1, BT2.1, BTF1 | 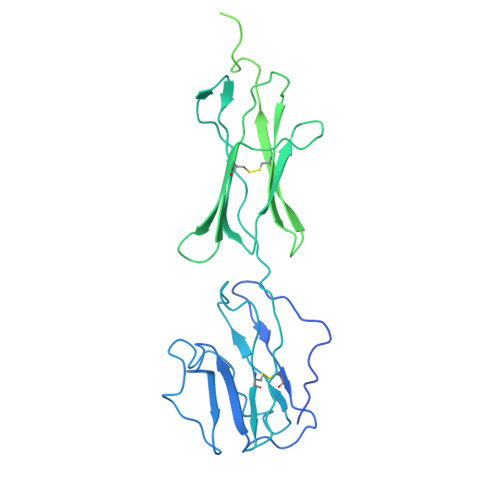 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q7KYR7 GTEx: ENSG00000112763 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q7KYR7 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Butyrophilin subfamily 3 member A2 | C [auth E] | 340 | Homo sapiens | Mutation(s): 0 Gene Names: BTN3A2, BT3.2, BTF3, BTF4 | 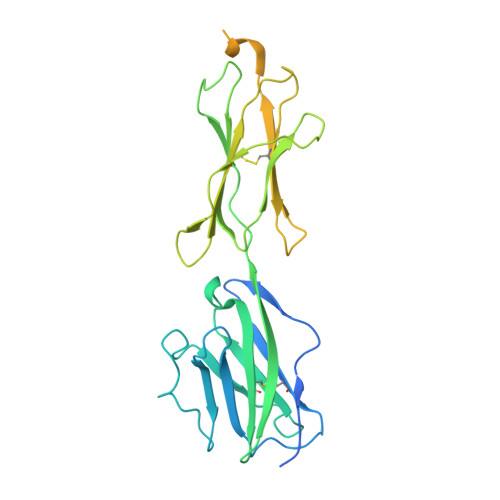 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P78410 GTEx: ENSG00000186470 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P78410 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Butyrophilin subfamily 3 member A1 | D [auth F] | 539 | Homo sapiens | Mutation(s): 0 Gene Names: BTN3A1, BTF5 | 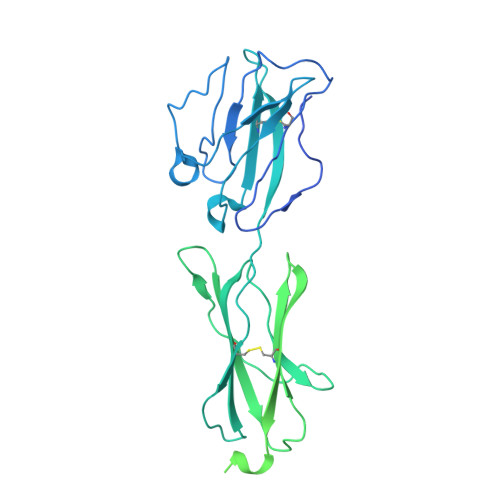 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O00481 GTEx: ENSG00000026950 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O00481 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ICT01 heavy chain | E [auth H], G [auth h] | 255 | Homo sapiens | Mutation(s): 0 | 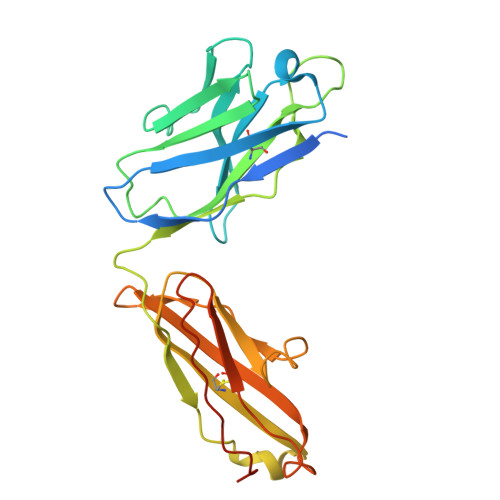 |
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ICT01 light chain | F [auth L], H [auth l] | 236 | Homo sapiens | Mutation(s): 0 |  |
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | 1.20.1 |
| RECONSTRUCTION | RELION | 5 |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | -- |