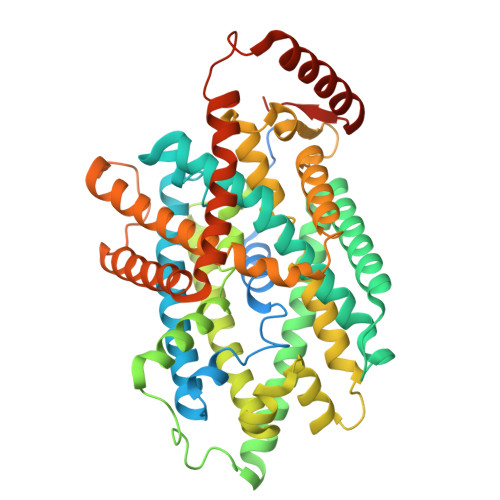
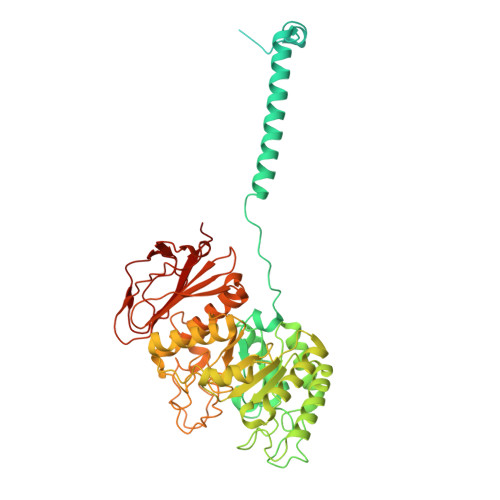
Structural insights into the human system y + L amino acid transporter complex.
Dai, L., Xu, K., Zhang, T., Wang, X., Zeng, Q., Liang, H., Xu, C., Yang, L., Wang, Z., Yan, R.(2025) Structure 33: 1224-1232.e5
- PubMed: 40347950 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2025.04.012
- Primary Citation Related Structures:
9KY5, 9LDR - PubMed Abstract:
System y + L facilitates the sodium-independent transport of cationic and sodium-dependent transport of neutral amino acids via heteromeric amino acid transporters. System y + L consists of either SLC7A6 (y + LAT2) or SLC7A7 (y + LAT1) and 4F2hc (SLC3A2). The y + LAT2-4F2hc complex mediates the exchange of L -lysine (Lys), L -arginine (Arg), L -leucine (Leu), and L -glutamine (Gln) and is important for the glutamate-glutamine cycle and ammonia clearance. c-Myc-driven upregulation of y + LAT2 in cancer enhances amino acid uptake and mTORC1 activation, promoting tumor growth. Its transport mechanism has remained unclear. Here, we determined the cryoelectron microscopic (cryo-EM) structures of the y + LAT2-4F2hc complex bound to either Arg or Leu at 3.60 Å and 3.58 Å resolution, respectively, revealing an outward-open conformation. Our structural analysis highlights conformational changes during transport, and functional assays validate critical residues involved in substrate binding and transport. These findings elucidate the molecular mechanism of the system y + L and provide a foundation for developing targeted therapies against y + LAT2.
- SUSTech Homeostatic Medicine Institute, School of Medicine, Southern University of Science and Technology, Shenzhen, Guangdong Province, China; Key University Laboratory of Metabolism and Health of Guangdong, Institute for Biological Electron Microscopy, Southern University of Science and Technology, Shenzhen, Guangdong Province, China.
Organizational Affiliation: