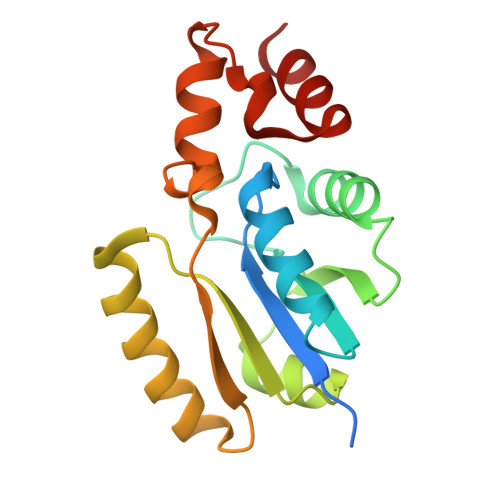
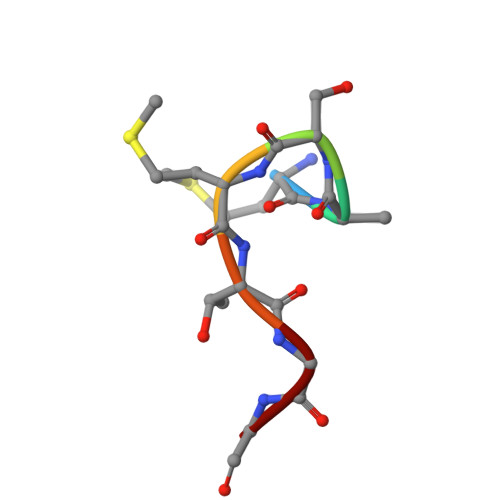
Protein-tags and their fragments as potent inhibitors of enzymes: Structure of the ternary complex of phosphopantetheine adenylyltransferase from Enterobacter spp. with tag-peptides and phosphonoacetic acid at 2.20 angstrom resolution.
Ahmad, N., Kumar, V., Goel, V.K., Sharma, P., Sharma, S., Singh, T.P.(2025) Protein Sci 34: e70216-e70216
- PubMed: 40689715 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.70216
- Primary Citation Related Structures:
9KPY - PubMed Abstract:
Phosphopantetheine adenylyltransferase (PPAT) catalyzes the penultimate step of coenzyme A (CoA) biosynthesis pathway by transferring the adenylyl group from adenosine triphosphate (ATP) to 4'-phosphopantetheine (PNS), yielding 3'-dephosphocoenzyme A and pyrophosphate. In this study, the recombinant PPAT from Enterobacter spp. strain 638 (EbPPAT) was purified and co-crystallized with phosphonoacetic acid (PAE). The structure showed the presence of three homodimers AB, CD, and EF in the asymmetric unit. The 14 extra N-terminal residues (Met-14 to Ser-1, 14-mer peptide) from the expression tag were observed in Molecules B and F. These tag-peptides occupied the PNS-binding sites of adjacent Molecules A and E, respectively. Additionally, a heptapeptide (Met-14 to Gly-8) was also observed in the PNS-binding site of Molecule C. Furthermore, two PAE molecules were present in the ATP-binding sites of Molecules B, D, and F, whereas a single PAE molecule was found in Molecules A, C, and E. This showed that tag-peptides blocked the PNS-binding site while PAE blocked the ATP-binding sites. Three peptides of the tag, including 14-mer (Met-14 to Ser-1), heptapeptide (Met-14 to Gly-8) and pentapeptide (Met-14 to Thr-10) were synthesized, and their binding affinities were estimated, which showed the K D values of 5.5 × 10 -5 , 1.8 × 10 -8 , and 7.3 × 10 -9 M, respectively. PAE molecules bound to EbPPAT in the ATP-binding sites with a K D of 4.77 × 10 -4 M. This is the first structure of PPAT with peptides bound in the substrate-binding sites, indicating a novel approach to design peptide inhibitors.
- Department of Biophysics, All India Institute of Medical Sciences, New Delhi, India.
Organizational Affiliation: