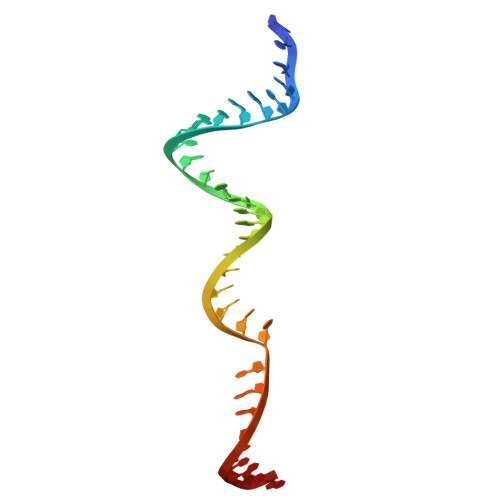
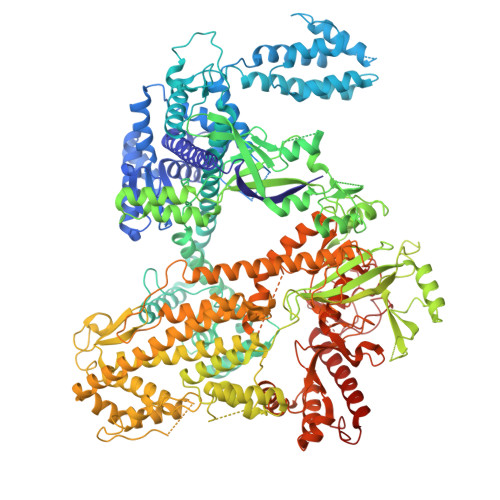
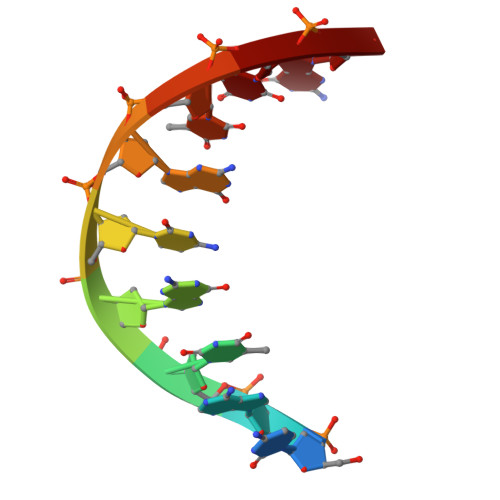
Catalytic-state structure of Candidatus Hydrogenedentes Cas12b revealed by cryo-EM studies.
Li, Y., Li, J., Pei, X., Wei, J., Gan, J., Lin, J.(2025) Nucleic Acids Res 53
- PubMed: 40539514 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaf519
- Primary Citation Related Structures:
9KLN, 9KLO, 9KLP, 9KLQ - PubMed Abstract:
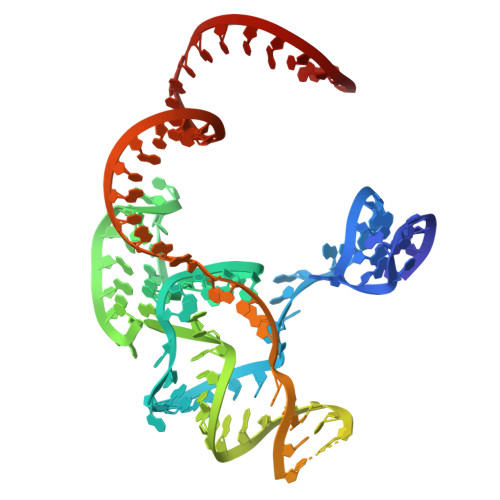
The CRISPR-Cas (clustered regularly interspaced short palindromic repeats and CRISPR-associated protein) systems are adaptive immune mechanisms that play critical roles in defending archaea and bacteria against invading entities. These systems can be divided into two classes, with class 2 comprising three types (II, V, and VI). Because of their ability to cleave double-stranded DNA, many class 2 CRISPR-Cas proteins have been harnessed as genome editing tools. Unlike the well-studied type II Cas9 proteins, the structural studies of the type V-B Cas12b proteins are limited, hindering their engineering and broader application. Here, we report four complex structures of ChCas12b, which reveal many unique structural features. The folding of the single guide RNA (sgRNA) of ChCas12b is distinct from that of AacCas12b and BthCas12b. Notably, many of these unique features are involved in ChCas12b-sgRNA interaction, suggesting that they are co-evolved. While ChCas12b shares a conserved two-cation-assisted catalytic mechanism with its homologs, it recognizes a longer guide:target heteroduplex, potentially offering higher fidelity in DNA editing. Altogether, our studies suggested that Cas12b family proteins exhibit significant diversity in their folding, sgRNA and target DNA binding. In the future, it is worth characterizing more representative proteins to identify CRISPR-Cas proteins with higher gene editing ability and fidelity.
- State Key Laboratory of Genetics and Development of Complex Phenotypes, School of Life Sciences, Zhongshan Hospital, Fudan University, Shanghai 200438, China.
Organizational Affiliation: