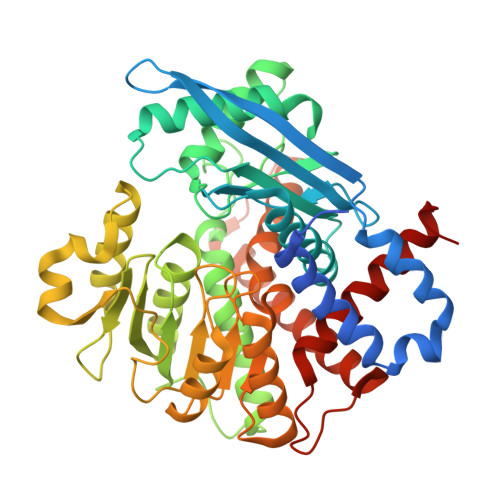
Phosphorylation-mediated regulation of the NADPH-dependent glutamate dehydrogenase, SpGdh1, from Schizosaccharomyces pombe.
Wang, Y.F., Tomita, T., Yoshida, A., Kosono, S., Nishiyama, M.(2025) J Biological Chem 301: 110422-110422
- PubMed: 40578557 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2025.110422
- Primary Citation Related Structures:
9KL6 - PubMed Abstract:
Glutamate dehydrogenase from the yeast Schizosaccharomyces pombe (SpGdh1) is a pivotal enzyme that catalyzes the conversion of 2-oxoglutarate and ammonium to glutamate using NADPH as a coenzyme. Although SpGdh1 is phosphorylated at several residues, the impact of phosphorylation on enzyme activity and the underlying molecular mechanisms remains unclear. To elucidate the phosphorylation-mediated regulation of SpGdh1, we determined the crystal structure of SpGdh1 binding 2-iminoglutarate (2-IG) and NADP + . The results of the structural analysis revealed that four serine residues for phosphorylation were located near the active site. Ser252 directly interacted with the 2'-phosphate group of the adenine ribose moiety of NADP + , suggesting that the phosphorylation of Ser252 interfered with NADP + binding. To confirm this hypothesis, we prepared SpGdh1 phosphorylation-mimic (Ser to Glu) variants of SpGdh1 at these four Ser residues. The results of a kinetic analysis revealed that the replacement of these four residues increased the apparent K m NADP(H) value and decreased catalytic efficiency, k cat /K m NADP(H) .In contrast, substitutions decreased the apparent K m NAD(H) value and increased catalytic efficiency, k cat /K m NAD(H) . Therefore, the Ser to Glu replacement caused net shifts in the coenzyme specificities (NADPH to NADH and NADP + to NAD + ) of 55- and 2900-fold, respectively. This is the first study to reveal the effects of the phosphorylation of SpGdh1 on its activity.
- Graduate School of Agricultural and Life Sciences, The University of Tokyo, Bunkyo-ku, Tokyo, Japan.
Organizational Affiliation: