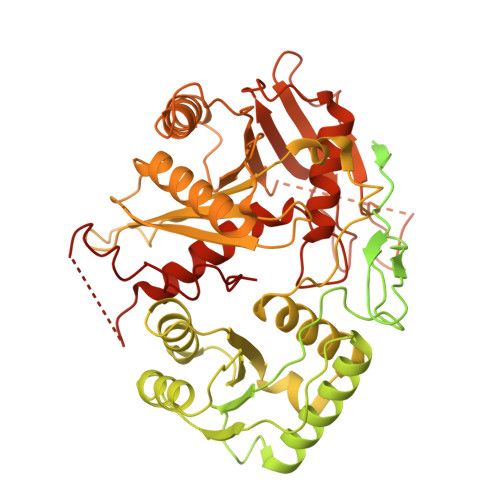
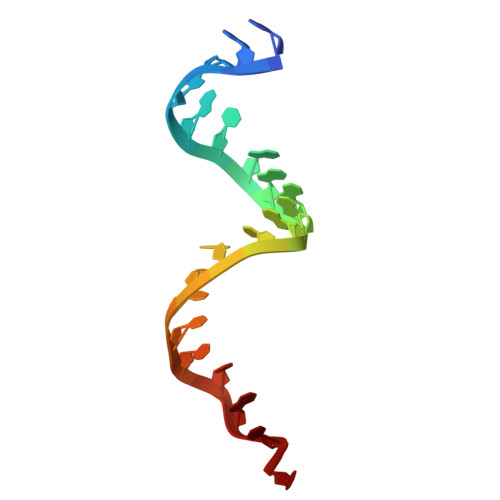
Mechanistic insights into RNA cleavage by human Argonaute2-siRNA complex.
Li, Z., Xu, Q., Zhang, Y., Zhong, J., Zhang, T., Xue, J., Liu, S., Gao, H., Zhang, Z.Z.Z., Wu, J., Shen, E.Z.(2025) Cell Res 35: 453-464
- PubMed: 40240484 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-025-01114-7
- Primary Citation Related Structures:
9K6P, 9K6Q, 9K6R, 9K6S, 9K6T - PubMed Abstract:
In animals, AGO-clade Argonaute proteins utilize small interfering RNAs (siRNAs) as guides to recognize target with complete complementarity, resulting in target RNA cleavage that is a critical step for target silencing. These proteins feature a constricted nucleic acid-binding channel that limits base pairing between the guide and target beyond the seed region. How the AGO-siRNA complexes overcome this structural limitation and achieve efficient target cleavage remains unclear. We performed cryo-electron microscopy of human AGO-siRNA complexes bound to target RNAs of increasing lengths to examine the conformational changes associated with target recognition and cleavage. Initially, conformational transition propagates from the opening of the PAZ domain and extends through a repositioning of the PIWI-L1-N domain toward the binding channel, facilitating the capture of siRNA-target duplex. Subsequent extension of base pairing drives the downward movement of the PIWI-L1-N domain to enable catalytic activation. Finally, further base pairing toward the 3' end of siRNA destabilizes the PAZ-N domain, resulting in a "uni-lobed" architecture, which might facilitate the multi-turnover action of the AGO-siRNA enzyme complex. In contrast to PIWI-clade Argonautes, the "uni-lobed" structure of the AGO complex makes multiple contacts with the target in the central region of the siRNA-target duplex, positioning it within the catalytic site. Our findings shed light on the stepwise mechanisms by which the AGO-siRNA complex executes target RNA cleavage and offer insights into the distinct operational modalities of AGO and PIWI proteins in achieving such cleavage.
- Fudan University, Shanghai, China.
Organizational Affiliation: