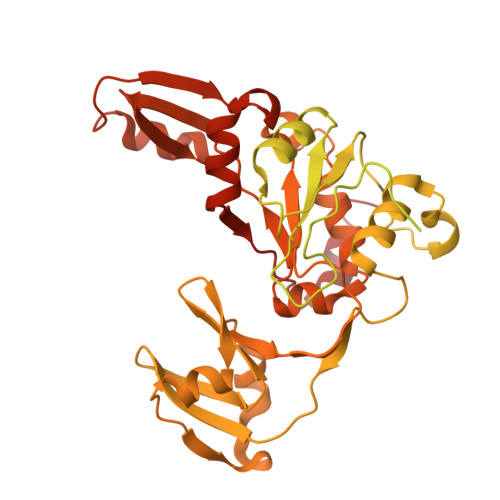
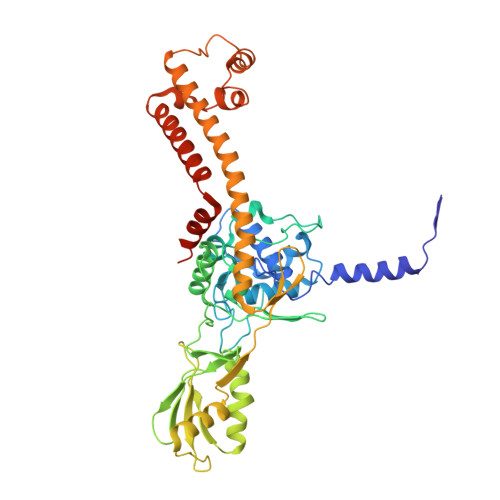
Direct trapping of the transport-segment DNA by the central domain of type IIA topoisomerases.
Xin, Y., Xian, R., Liu, C., Zhang, O., Rao, Z., Li, X., Chen, Y.(2025) Sci Adv 11: eadw2839-eadw2839
- PubMed: 41406217 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adw2839
- Primary Citation Related Structures:
9JNK, 9JOR, 9LII - PubMed Abstract:
Type IIA topoisomerases modulate DNA topology by coordinating the cleavage of gate-segment DNA and the passage of transport-segment DNA-a mechanism conserved across species and essential for diverse cellular processes. While gate-segment interactions have been extensively studied, direct structural evidence of transport-segment capture has remained elusive, limiting our understanding of the full catalytic cycle. Here, we present a cryo-electron microscopy structure of the T4 bacteriophage topoisomerase II with a transport-segment DNA bound directly to its central domain. The structure reveals conformational rearrangements in the central domain that accommodate the transport-segment DNA, suggesting an alternative sequence of events in which the enzyme sliding along the loosely bound religated gate segment may precede transport-segment passage through the coiled-coil gate. Supported by mutational and biochemical assays, our findings provide previously unidentified mechanistic insights and open potential avenues for the development of next-generation type IIA topoisomerase inhibitors.
- State Key Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: