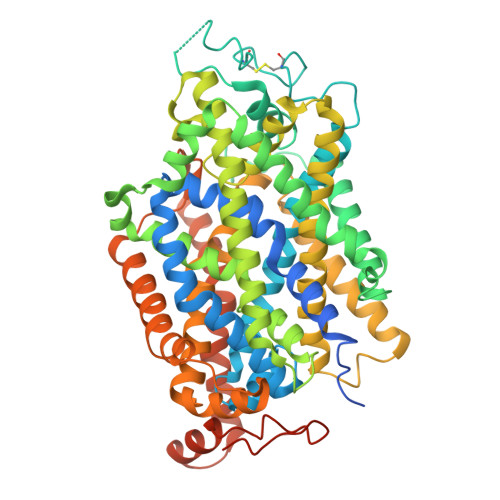
Dimerization and substrate recognition of human taurine transporter.
Zhang, Y., Chen, J., Chen, N., Xiong, H., Zhu, Z., Yang, D., Ge, J., Yu, J.(2025) Nat Commun 16: 6163-6163
- PubMed: 40615403 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-60967-z
- Primary Citation Related Structures:
9JD3, 9JD4, 9JD9, 9JDA, 9JDG, 9JDJ, 9JDL - PubMed Abstract:
Taurine is a conditionally essential nutrient and one of the most abundant amino acids in humans, with diverse physiological functions. The cellular uptake of taurine is primarily mediated by the taurine transporter (TauT), and its dysfunction leads to retinal regeneration, cardiomyopathy, neurological and aging-associated disorders. Here we determine structures of TauT in two states: the apo inward-facing open state and the occluded state bound with substrate taurine or γ-aminobutyric acid (GABA). In addition to monomer, the structures also reveal a TauT dimer, where two cholesterol molecules act as "molecular glue", and close contacts of two TM5 from each protomer mediate the dimer interface. In combination with functional characterizations, our results elucidate the detailed mechanisms of substrate recognition, specificity and transport by TauT, providing a structural framework for understanding TauT function and exploring potential therapeutic strategies for taurine-deficiency-related disorders.
- Interdisciplinary Research Center on Biology and Chemistry, Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: