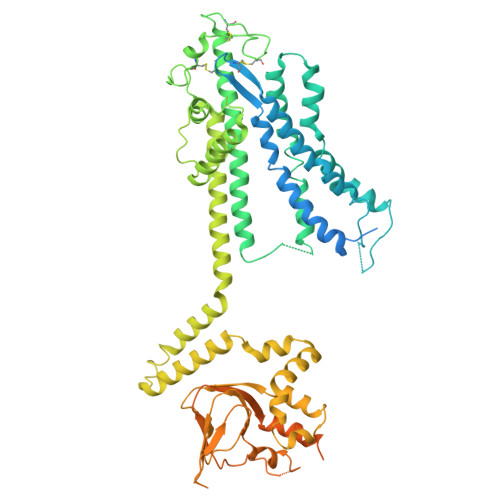
Cryo-EM structures of Arabidopsis CNGC1 and CNGC5 reveal molecular mechanisms underlying gating and calcium selectivity.
Wang, J., Du, B.Y., Zhang, X., Qu, X., Yang, Y., Yang, Z., Wang, Y.F., Zhang, P.(2025) Nat Plants 11: 632-642
- PubMed: 39979428 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41477-025-01923-z
- Primary Citation Related Structures:
9J34, 9J35, 9J36 - PubMed Abstract:
Plant cyclic nucleotide-gated channels (CNGCs) belong to the cyclic nucleotide-binding domain (CNBD) channel family, but are phylogenetically classified in a distinct branch. In contrast to their animal counterparts of K + -selective or non-selective cation channels, plant CNGCs mainly mediate Ca 2+ influx and are involved in various physiological processes, such as stomatal movements, pollen-tube growth and immune responses. Here, we present the cryo-EM structure and electrophysiological analysis of plant CNGC representatives, Arabidopsis CNGC1 and CNGC5. We found that CNGC1 and CNGC5 contain a unique extracellular domain featuring disulfide bonds that is essential for channel gating via coupling of the voltage-sensing domain with the pore domain. The pore domain selectivity filter possesses a Gln residue at the constriction site that determines the Ca 2+ selectivity. Replacement of this Gln with Glu, typically observed in CNBD-type non-selective cation channels, could convert CNGC1 and CNGC5 from Ca 2+ -selective channels to non-selective cation channels permeable to Ca 2+ , Na + or K + . In addition, we found that the CNGC1 and CNGC5 CNBD homology domain contains intrinsic-ligand-like interactions, which may devoid the binding of cyclic nucleotides and lead to gating independent of cAMP or cGMP. This research not only provides a mechanistic understanding of plant CNGCs' function, but also adds to the comprehensive knowledge of the CNBD channels.
- National Key Laboratory of Plant Molecular Genetics, CAS Center for Excellence in Molecular Plant Sciences, Institute of Plant Physiology and Ecology, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: