CryoEM structure of a transmembrane protein
Ning, Y., Ge, J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Sodium-dependent serotonin transporter | 636 | Homo sapiens | Mutation(s): 0 Gene Names: SLC6A4, HTT, SERT | 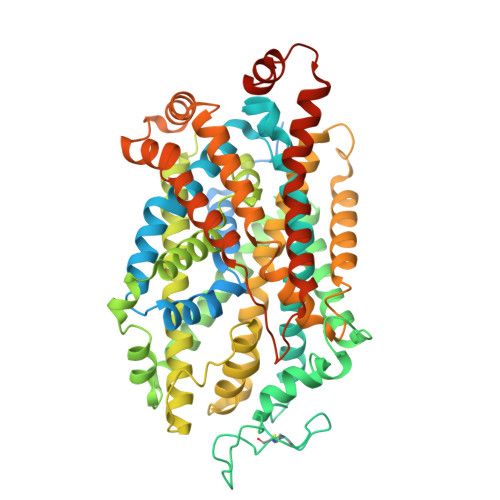 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P31645 GTEx: ENSG00000108576 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P31645 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: P31645-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 10 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| LMT Download:Ideal Coordinates CCD File | G [auth A] | DODECYL-BETA-D-MALTOSIDE C24 H46 O11 NLEBIOOXCVAHBD-QKMCSOCLSA-N |  | ||
| Y01 Download:Ideal Coordinates CCD File | E [auth A], F [auth A], W [auth A] | CHOLESTEROL HEMISUCCINATE C31 H50 O4 WLNARFZDISHUGS-MIXBDBMTSA-N |  | ||
| JC9 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | I [auth A] | (2~{S})-2-(2-chlorophenyl)-2-(methylamino)cyclohexan-1-one C13 H16 Cl N O YQEZLKZALYSWHR-ZDUSSCGKSA-N |  | ||
| NAG Download:Ideal Coordinates CCD File | H [auth A] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| D12 Download:Ideal Coordinates CCD File | AA [auth A] L [auth A] N [auth A] T [auth A] V [auth A] | DODECANE C12 H26 SNRUBQQJIBEYMU-UHFFFAOYSA-N |  | ||
| D10 Download:Ideal Coordinates CCD File | O [auth A], U [auth A] | DECANE C10 H22 DIOQZVSQGTUSAI-UHFFFAOYSA-N |  | ||
| DD9 Download:Ideal Coordinates CCD File | J [auth A], K [auth A], M [auth A], X [auth A], Z [auth A] | nonane C9 H20 BKIMMITUMNQMOS-UHFFFAOYSA-N |  | ||
| OCT Download:Ideal Coordinates CCD File | BA [auth A], P [auth A], Q [auth A], R [auth A], S [auth A] | N-OCTANE C8 H18 TVMXDCGIABBOFY-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | B [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| NA Download:Ideal Coordinates CCD File | C [auth A], D [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Funding Organization | Location | Grant Number |
|---|---|---|
| Other government | LG-QS-202203-05; 22PJ1410300 |