Neutralizing antibodies against Chikungunya virus and structural elucidation of their mechanism of action.
Han, X., Ji, C., Tian, S., Wang, F., Cao, G.P., Li, D., Duan, X., Tong, Z., Qi, J., Wang, Q., Huang, Q., Zhan, B.D., Gao, G.F., Yan, J.(2025) Nat Commun 16: 9682-9682
- PubMed: 41184282 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-64687-2
- Primary Citation Related Structures:
9IW2, 9IXA, 9IXI, 9IYI, 9J6D - PubMed Abstract:
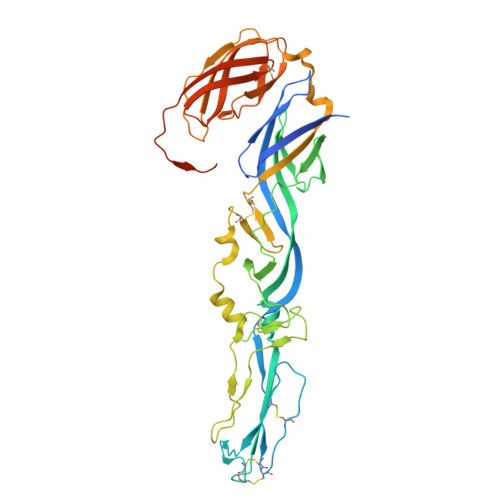
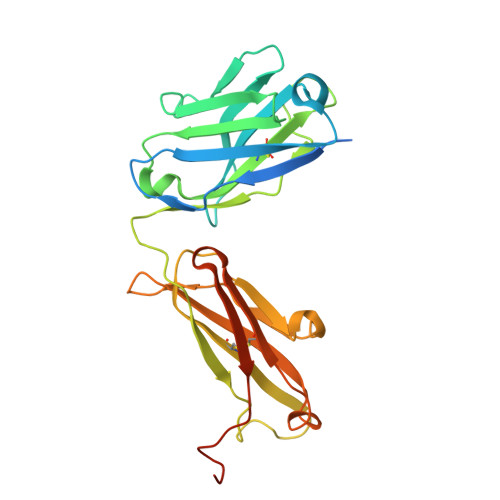
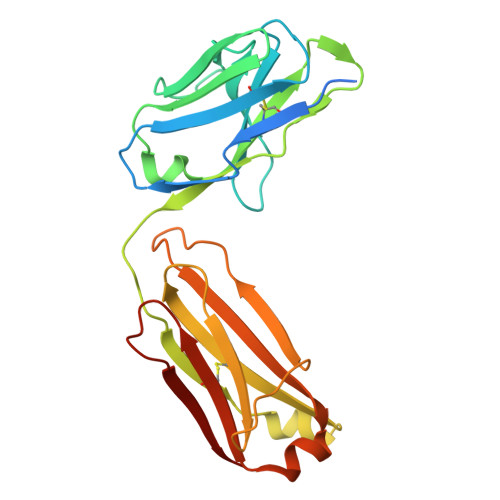
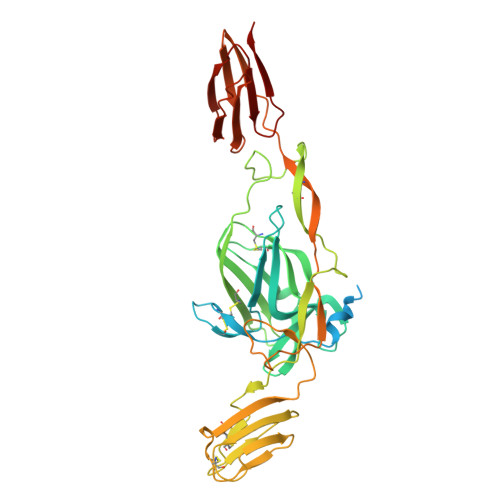
Chikungunya virus (CHIKV) is a mosquito-borne alphavirus that causes febrile illness and acute or chronic arthritis. Most therapeutics are still in the pre-clinical stage. In this study, we report the isolation of two neutralizing antibodies, C34 and C37, from a convalescent patient and investigate their mechanisms of action. Both C34 and C37 exhibit high neutralizing activities in vitro and demonstrate protective effects against CHIKV in a female mouse model. Our functional and structural studies reveal a mechanism that inhibits multiple stages of the virus infection cycle. Both antibodies bind with high affinity to an epitope spanning E2, E1, and the connecting β-strands, facilitating intra- and inter-virion crosslinking. Cryo-EM structures additionally identify a minor patch located beneath the E3 binding site on E2, which is allosterically exposed upon E3 dissociation during virus maturation. Functional and structural data further suggest that binding to the CHIKV receptor, Mxra8, is obstructed due to a clash between the antibodies and the stalk region of Mxra8. Our results highlight the potential of antibody-based therapeutics against CHIKV and elucidate the mechanisms of monoclonal antibody protection.
- CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: