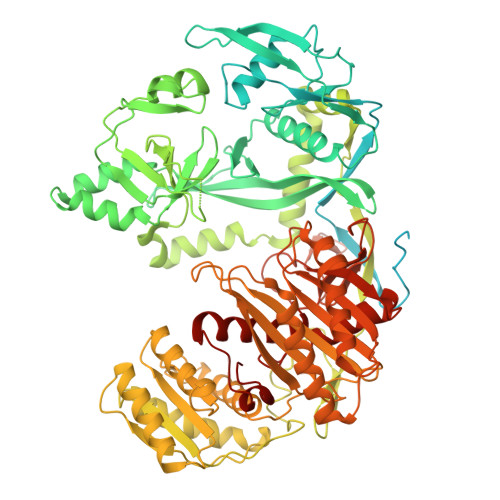
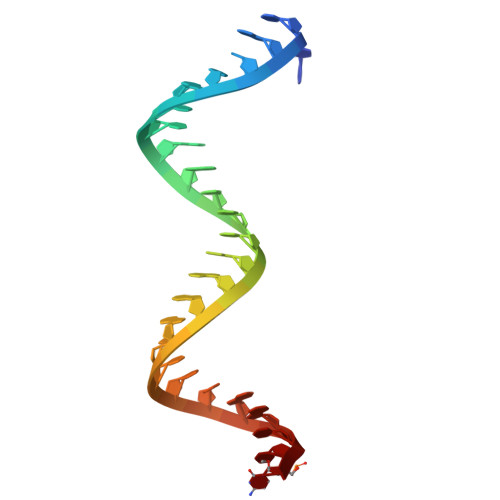
Structural insights into RNA cleavage by PIWI Argonaute.
Li, Z., Xu, Q., Zhong, J., Zhang, Y., Zhang, T., Ying, X., Lu, X., Li, X., Wan, L., Xue, J., Huang, J., Zhen, Y., Zhang, Z., Wu, J., Shen, E.Z.(2025) Nature 639: 250-259
- PubMed: 39814893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-08438-1
- Primary Citation Related Structures:
9IIY, 9IIZ, 9IJ0, 9IJ1, 9IJ2, 9IJ3, 9IJ4, 9IJ5 - PubMed Abstract:
Argonaute proteins are categorized into AGO and PIWI clades. Across most animal species, AGO-clade proteins are widely expressed in various cell types, and regulate normal gene expression 1 . By contrast, PIWI-clade proteins predominantly function during gametogenesis to suppress transposons and ensure fertility 1,2 . Both clades use nucleic acid guides for target recognition by means of base pairing, crucial for initiating target silencing, often through direct cleavage. AGO-clade proteins use a narrow channel to secure a tight guide-target interaction 3 . By contrast, PIWI proteins feature a wider channel that potentially allows mismatches during pairing, broadening target silencing capability 4,5 . However, the mechanism of PIWI-mediated target cleavage remains unclear. Here we demonstrate that after target binding, PIWI proteins undergo a conformational change from an 'open' state to a 'locked' state, facilitating base pairing and enhancing target cleavage efficiency. This transition involves narrowing of the binding channel and repositioning of the PIWI-interacting RNA-target duplex towards the MID-PIWI lobe, establishing extensive contacts for duplex stabilization. During this transition, we also identify an intermediate 'comma-shaped' conformation, which might recruit GTSF1, a known auxiliary protein that enhances PIWI cleavage activity 6 . GTSF1 facilitates the transition to the locked state by linking the PIWI domain to the RNA duplex, thereby expediting the conformational change critical for efficient target cleavage. These findings explain the molecular mechanisms underlying PIWI-PIWI-interacting RNA complex function in target RNA cleavage, providing insights into how dynamic conformational changes from PIWI proteins coordinate cofactors to safeguard gametogenesis.
- Key Laboratory of Growth Regulation and Translational Research of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou, China.
Organizational Affiliation: