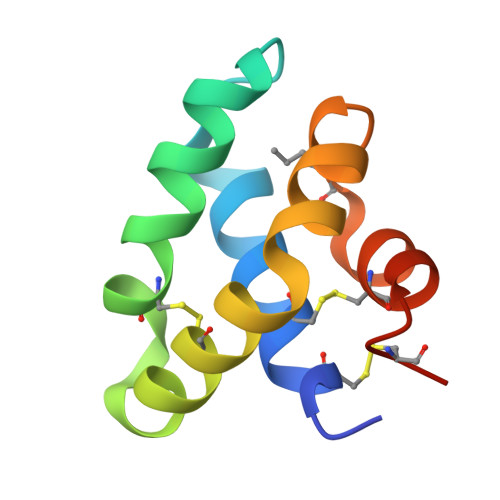
Synthetic Glycoforms Reveal Carbohydrate-Dependent Bioactivity of Human Saposin D.
Graf, C.G.F., Schulz, C., Schmalzlein, M., Heinlein, C., Monnich, M., Perkams, L., Puttner, M., Boos, I., Hessefort, M., Lombana Sanchez, J.N., Weyand, M., Steegborn, C., Breiden, B., Ross, K., Schwarzmann, G., Sandhoff, K., Unverzagt, C.(2017) Angew Chem Int Ed Engl 56: 5252-5257
- PubMed: 28378443 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201701362
- Primary Citation Related Structures:
9I63 - PubMed Abstract:
The main glycoforms of the hydrophobic lysosomal glycoprotein saposin D (SapD) were synthesized by native chemical ligation. An approach for the challenging solid-phase synthesis of the fragments was developed. Three SapD glycoforms were obtained following a general and robust refolding and purification protocol. A crystal structure of one glycoform confirmed its native structure and disulfide pattern. Functional assays revealed that the lipid-binding properties of three SapD glycoforms are highly affected by the single sugar moiety of SapD showing a dependency of the size and the type of N-glycan.
- Bioorg. Chemie, Gebäude NWI, Universität Bayreuth, 95440, Bayreuth, Germany.
Organizational Affiliation: