A Coiled Coil Module Strategy for High-Resolution Cryo-EM Structures of Small Proteins for Drug Discovery
Samson, C., Dossou, I., Steinmetz, A., Kumar, A., Mathieu, M., Rak, A.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Isoform 2B of GTPase KRas,APH2 | A, D [auth B] | 204 | Homo sapiens | Mutation(s): 1 Gene Names: KRAS, KRAS2, RASK2 EC: 3.6.5.2 | 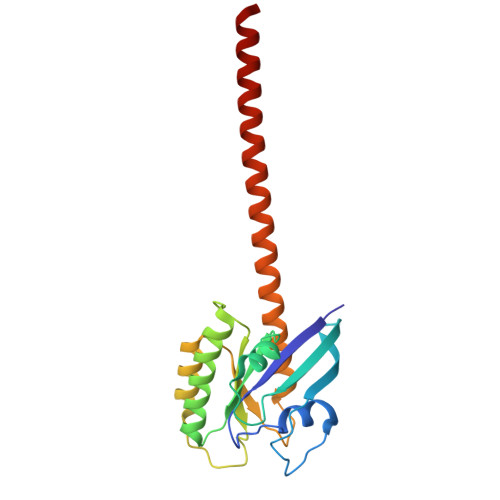 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P01116 GTEx: ENSG00000133703 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P01116 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Nanobody 26 | B [auth M], C [auth N] | 122 | Lama glama | Mutation(s): 0 | 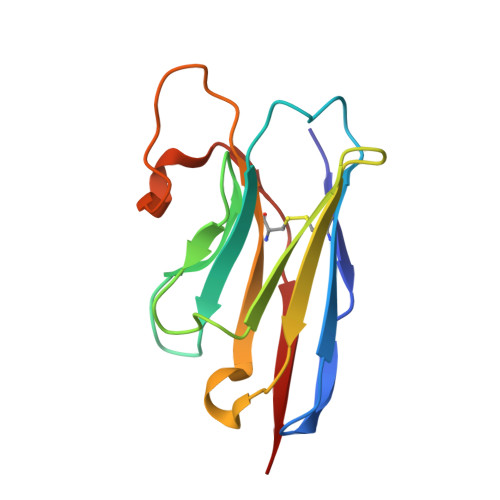 |
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| M1X Download:Ideal Coordinates CCD File | E [auth A], J [auth B] | {(2S)-4-[7-(8-chloronaphthalen-1-yl)-2-{[(2S)-1-methylpyrrolidin-2-yl]methoxy}-5,6,7,8-tetrahydropyrido[3,4-d]pyrimidin-4-yl]-1-[(2S)-2-fluoropropanoyl]piperazin-2-yl}acetonitrile C32 H37 Cl F N7 O2 BQFJNAAUAQEWHZ-XWGVYQGASA-N |  | ||
| GDP Download:Ideal Coordinates CCD File | F [auth A], H [auth B] | GUANOSINE-5'-DIPHOSPHATE C10 H15 N5 O11 P2 QGWNDRXFNXRZMB-UUOKFMHZSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | G [auth A], I [auth B] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | 1.21.2_5419 |
| RECONSTRUCTION | cryoSPARC |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | France | -- |