Cell cycle regulation has shaped replication origins in budding yeast.
Lim, C.T., Miller, T.C.R., Tan, K.W., Talele, S., Early, A., East, P., Sanchez, H., Dekker, N.H., Costa, A., Diffley, J.F.X.(2025) Nat Struct Mol Biol 32: 1697-1707
- PubMed: 40588661 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-025-01591-9
- Primary Citation Related Structures:
8RIF, 8RIG, 9I3I - PubMed Abstract:
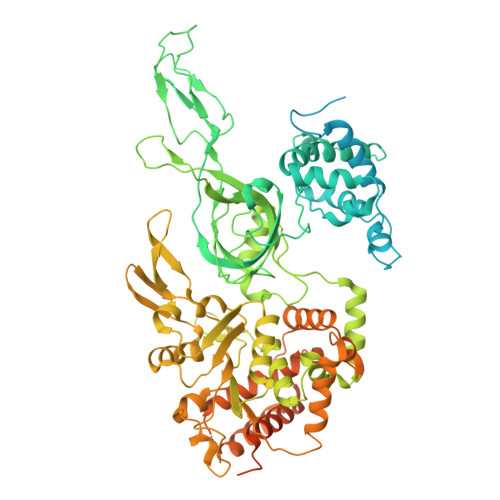
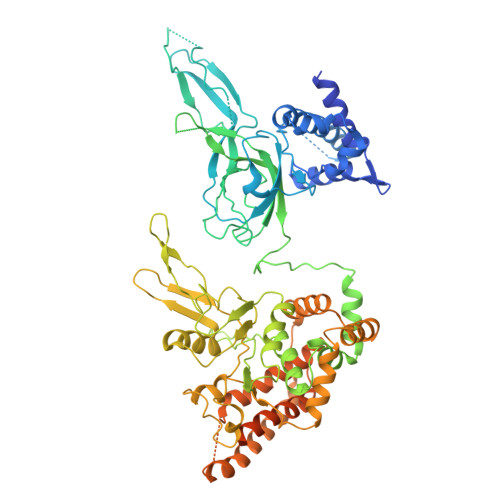
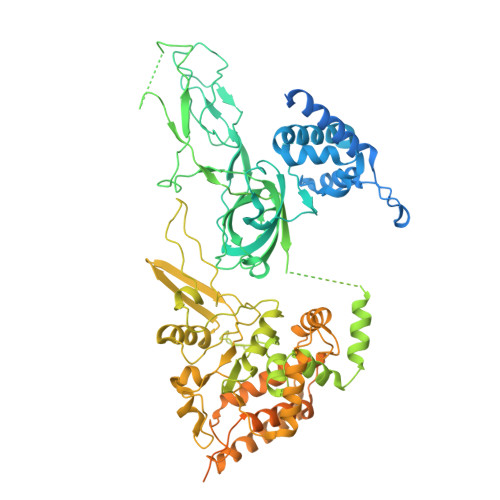
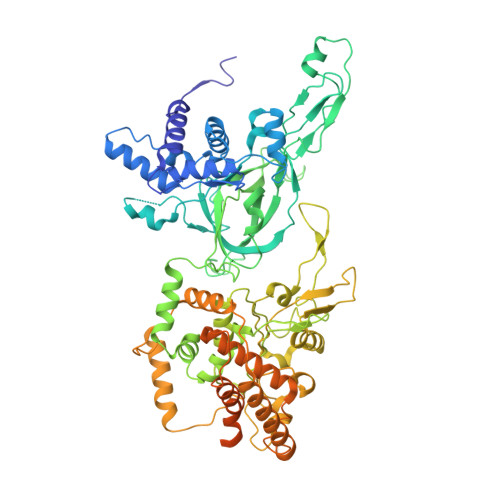
Eukaryotic DNA replication initiates from genomic loci known as origins. At budding yeast origins like ARS1, a double hexamer (DH) of the MCM replicative helicase is assembled by origin recognition complex (ORC), Cdc6 and Cdt1 by sequential hexamer loading from two opposed ORC binding sites. Cyclin-dependent kinase (CDK) inhibits DH assembly, which prevents re-replication by restricting helicase loading to the G1 phase. Here, we show that an intrinsically disordered region (IDR) in the Orc2 subunit promotes interaction between ORC and the first loaded, closed-ring MCM hexamer (the MCM-ORC (MO) intermediate). CDK-dependent phosphorylation of this IDR blocks MO formation and DH assembly. We show that MO stabilizes ORC at lower-affinity binding sites required for second hexamer loading. Origins comprising two high-affinity ORC sites can assemble DH efficiently without MO by independently loading single hexamers. Strikingly, these origins escape CDK inhibition in vitro and in vivo. Our work reveals mechanistic plasticity in MCM loading with implications for understanding how CDK regulation has shaped yeast origin evolution and how natural, strong origins might escape cell cycle regulation. We also identify key steps common to loading pathways, with implications for understanding how MCM is loaded in other eukaryotes.
- Chromosome Replication Laboratory, The Francis Crick Institute, London, UK.
Organizational Affiliation: