Cryo-EM structure of human sortilin ectodomain in complex with a thyroglobulin C-terminal peptide
Boniardi, I., Coscia, F.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Sortilin | 742 | Homo sapiens | Mutation(s): 0 Gene Names: SORT1 |  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q99523 GTEx: ENSG00000134243 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q99523 | ||||
Glycosylation | |||||
| Glycosylation Sites: 3 | Go to GlyGen: Q99523-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Thyroglobulin | 20 | Homo sapiens | Mutation(s): 0 |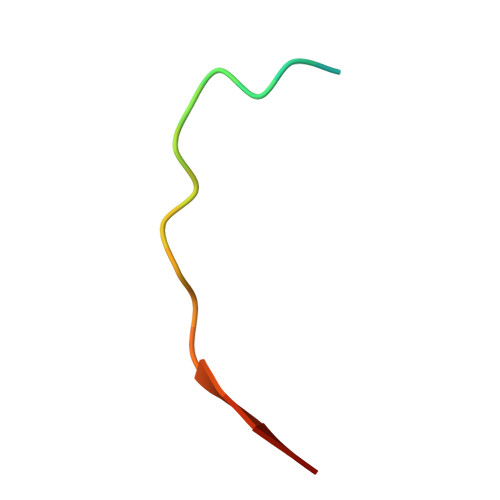  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P01266 GTEx: ENSG00000042832 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P01266 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | E [auth A] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | 1.21.1_5286 |
| Funding Organization | Location | Grant Number |
|---|---|---|
| European Research Council (ERC) | European Union | 101041298 |