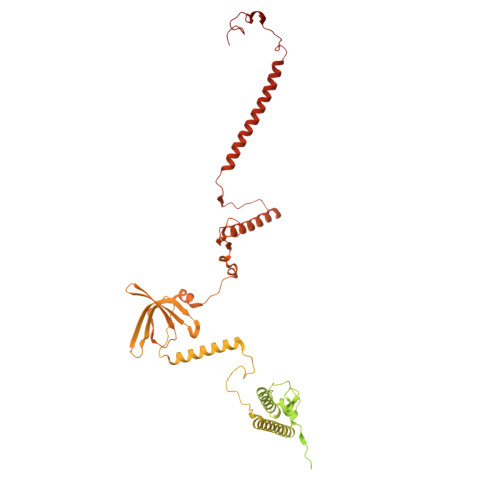
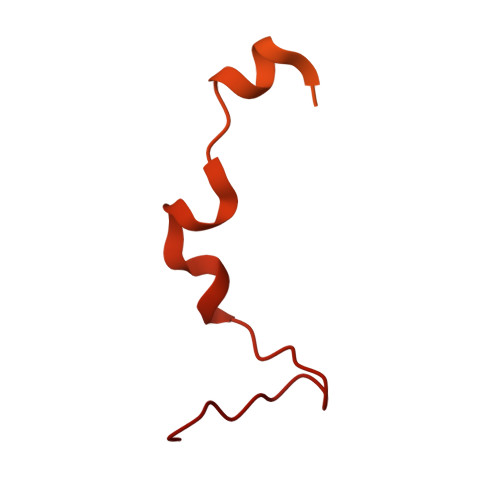
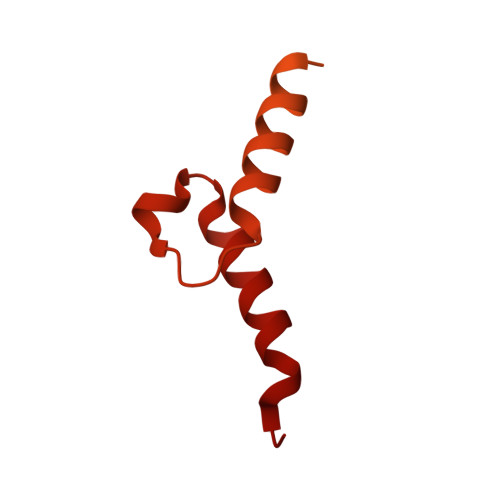
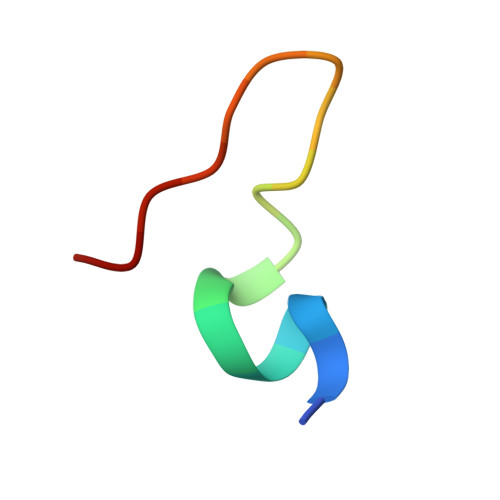
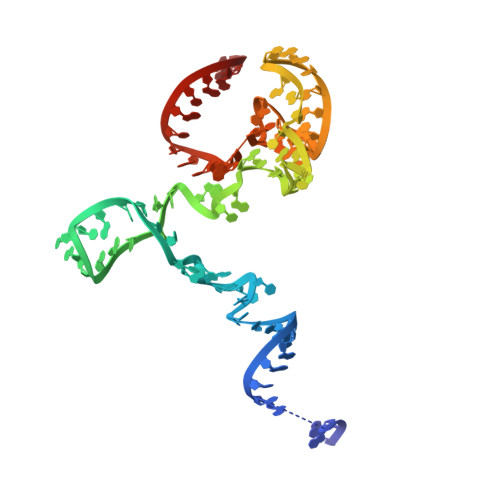
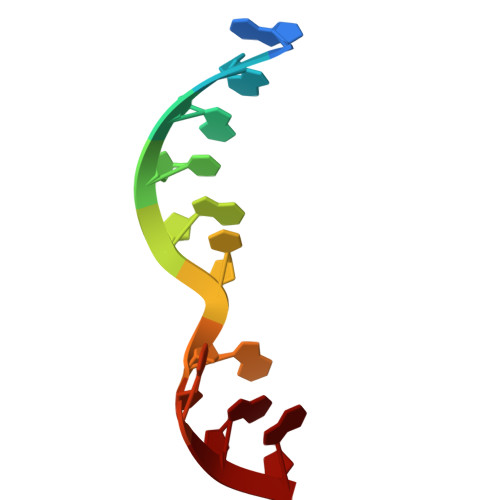
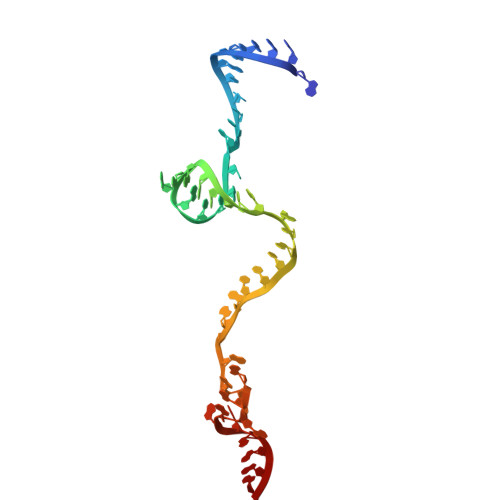
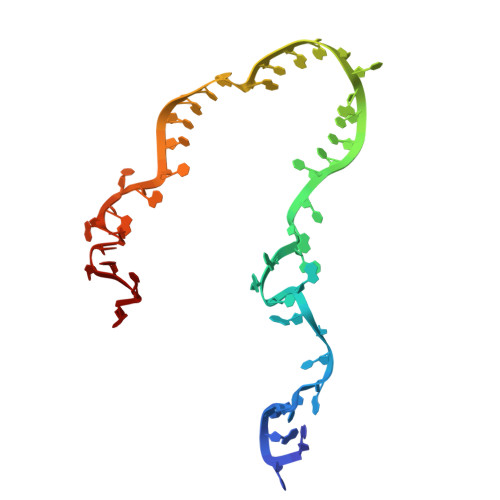
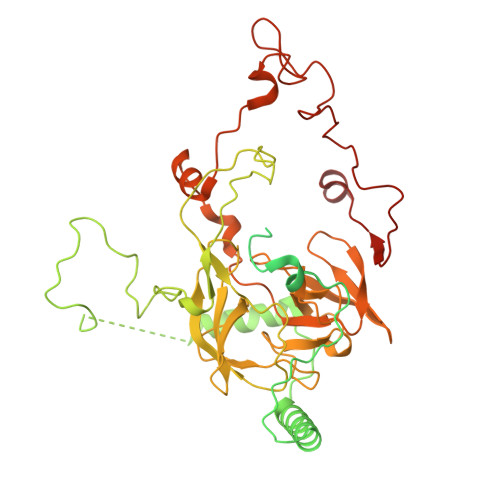
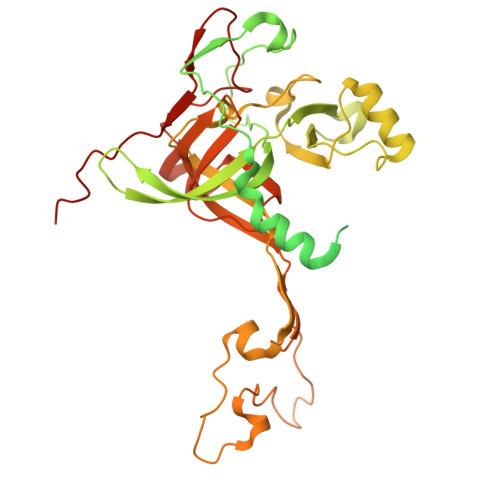
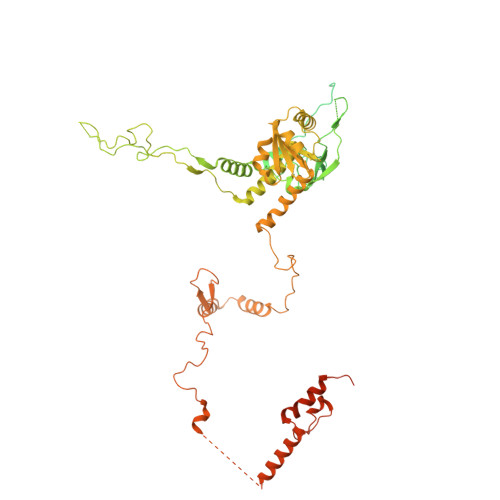
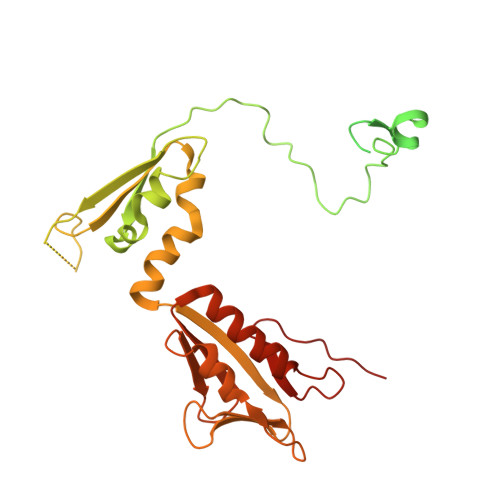
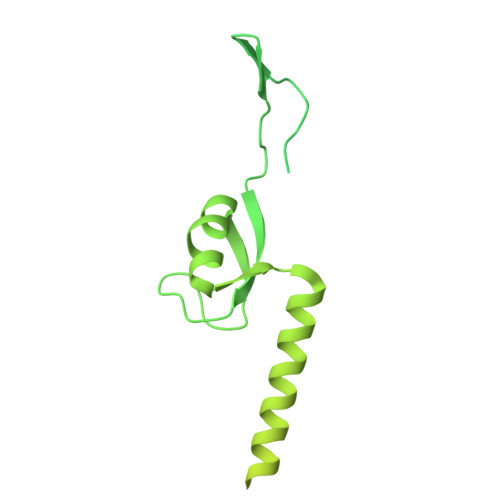
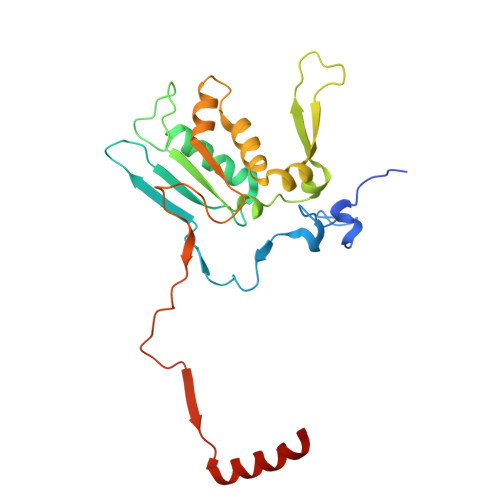
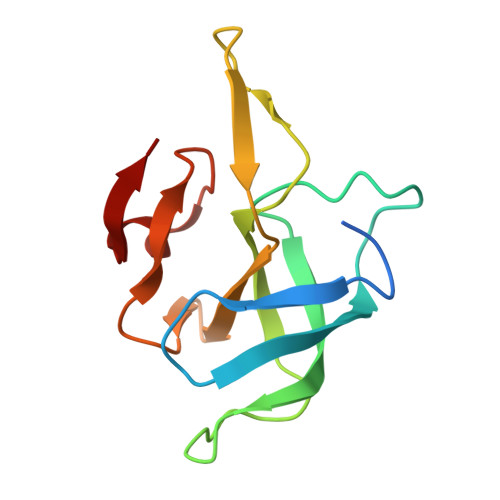
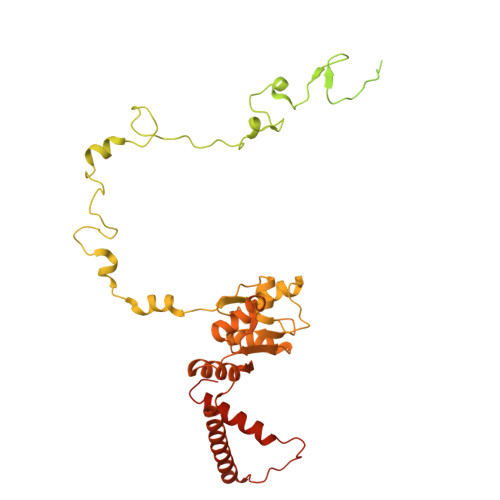
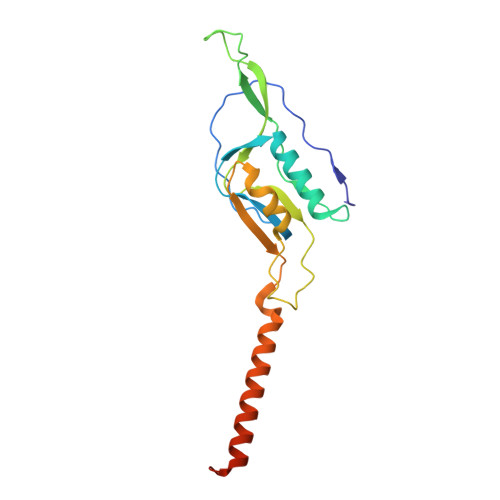
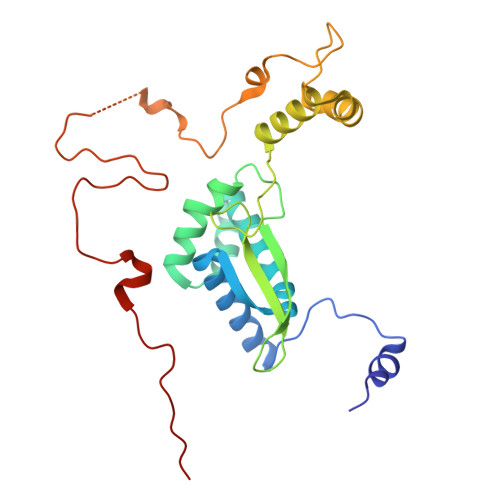
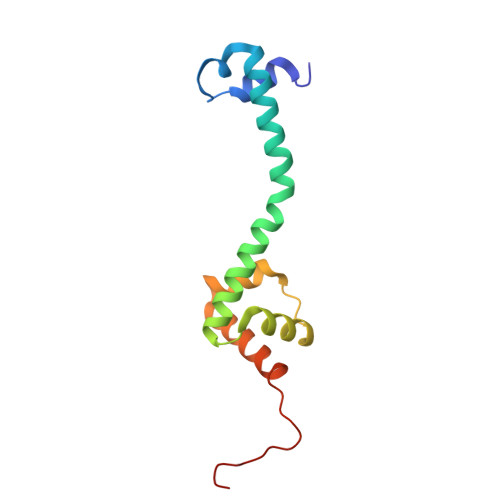
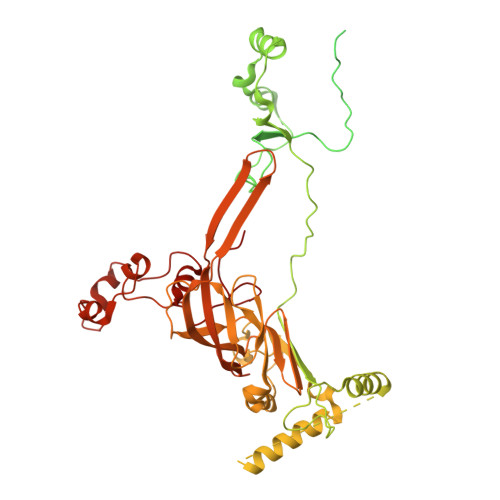
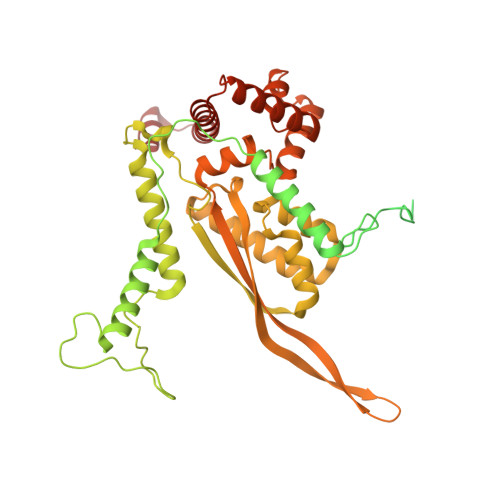
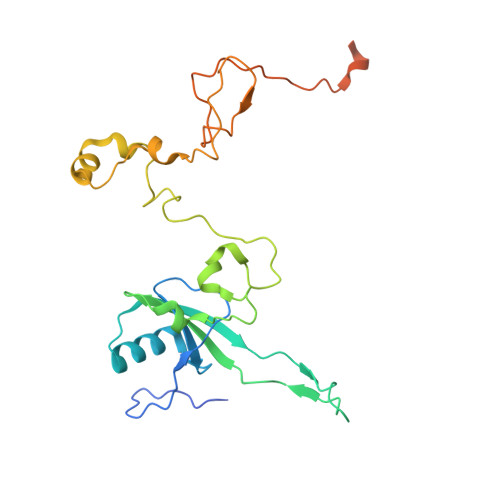
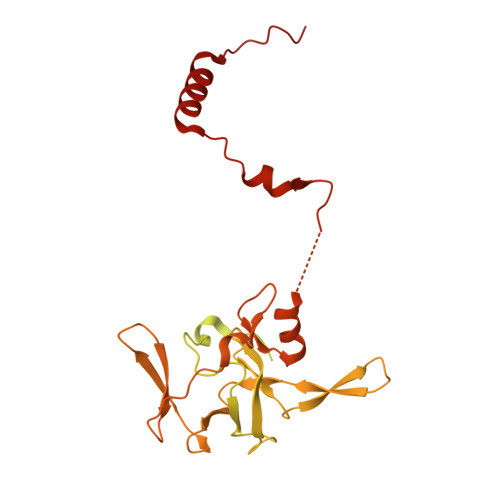
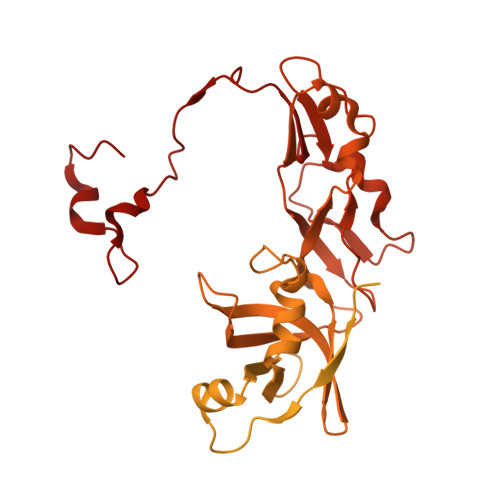
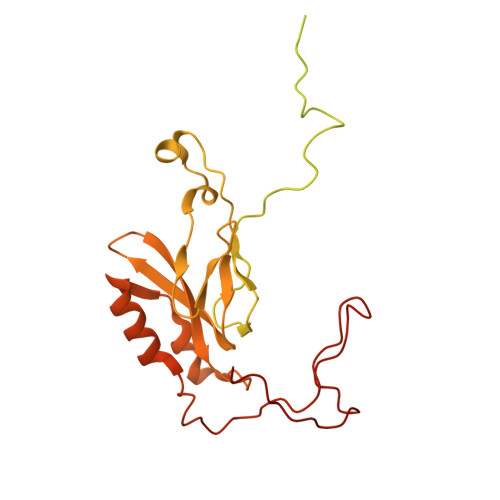
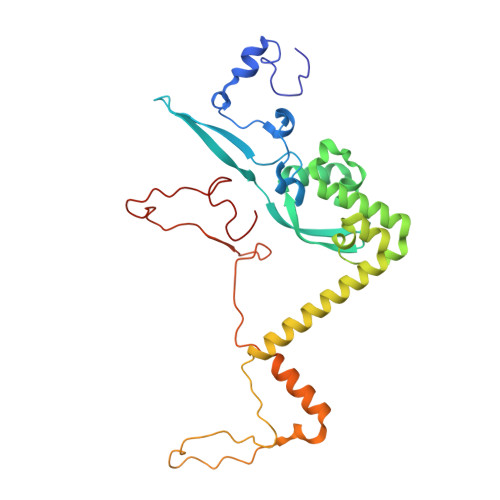
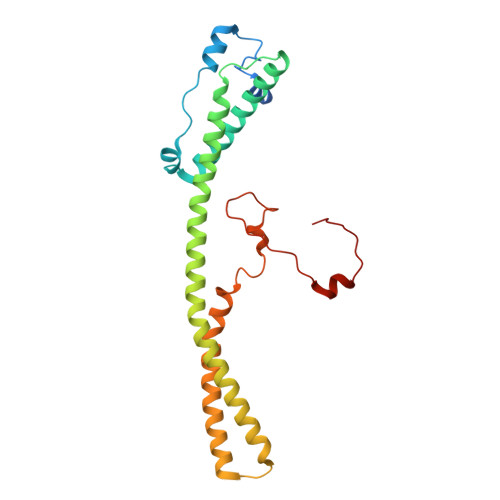
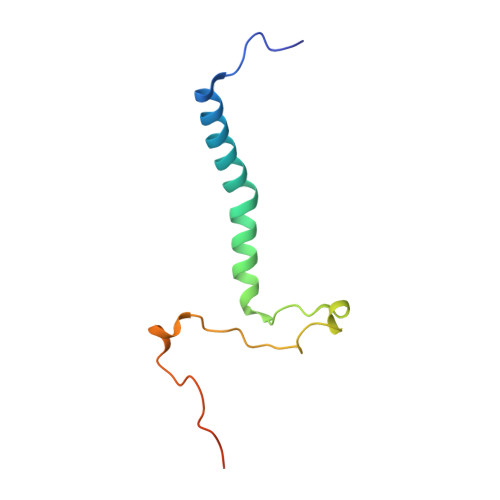
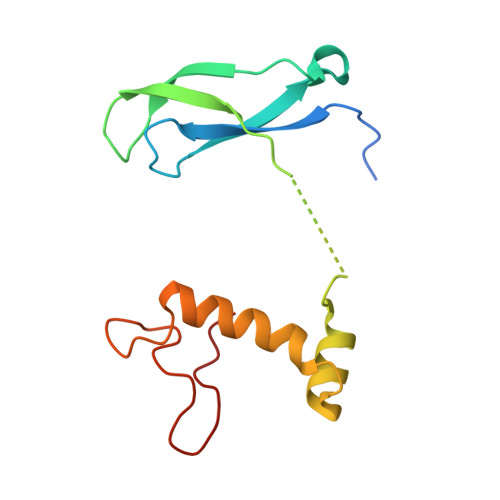
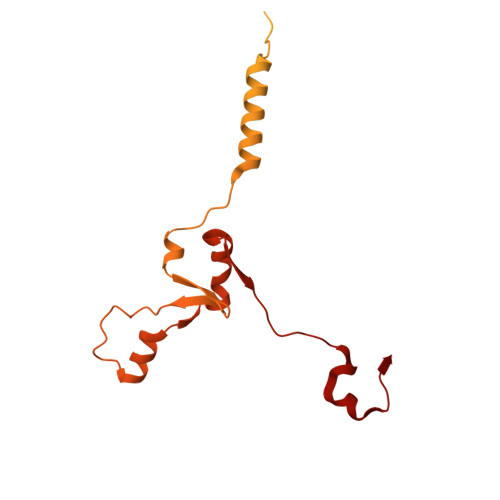
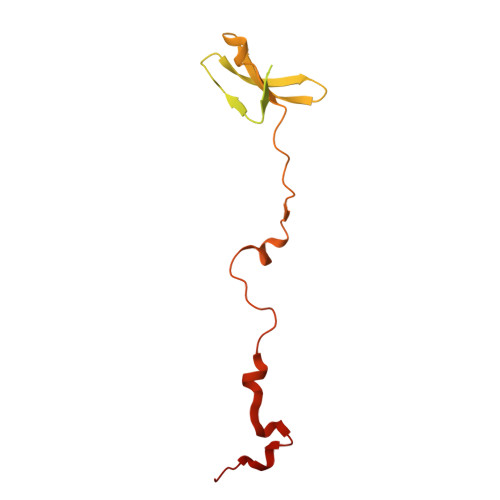
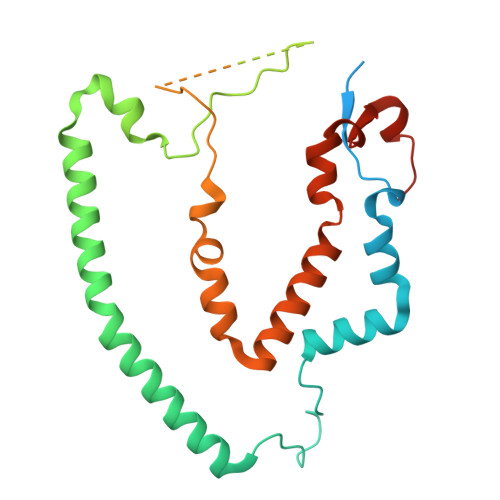
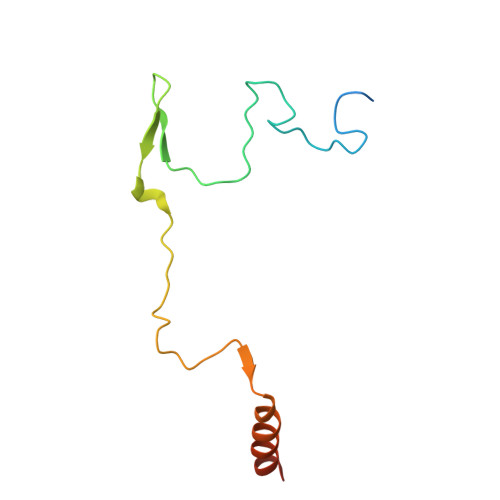
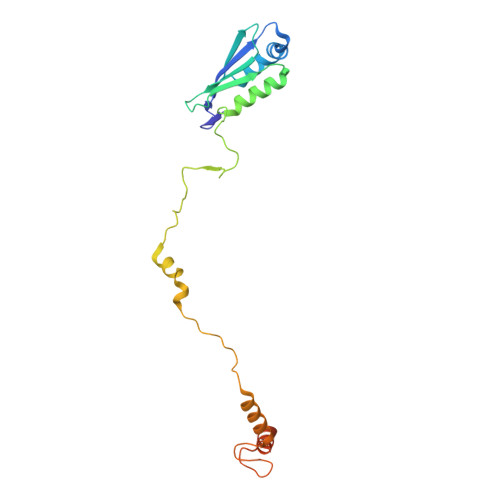
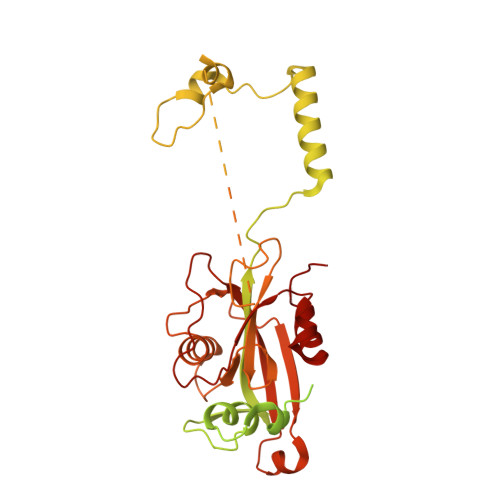
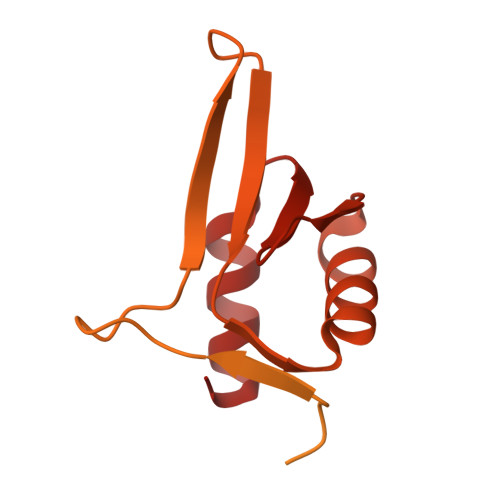
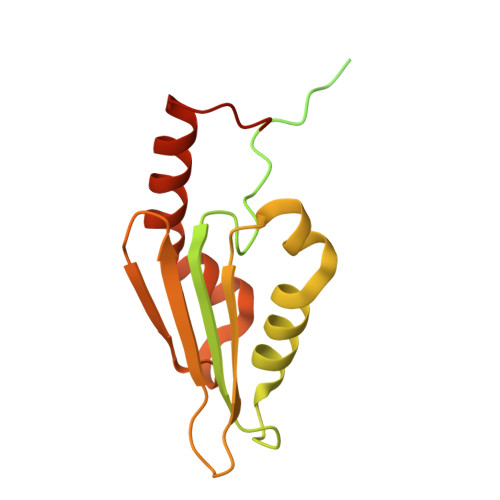
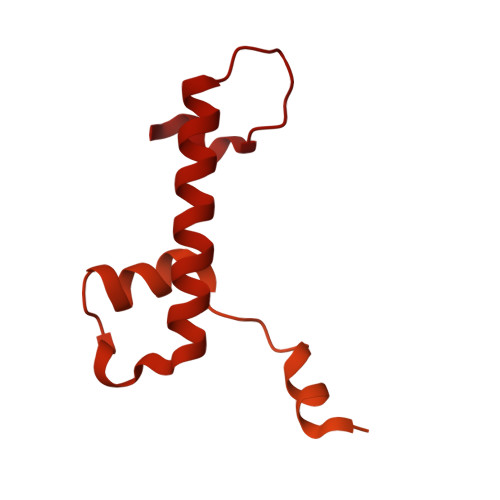
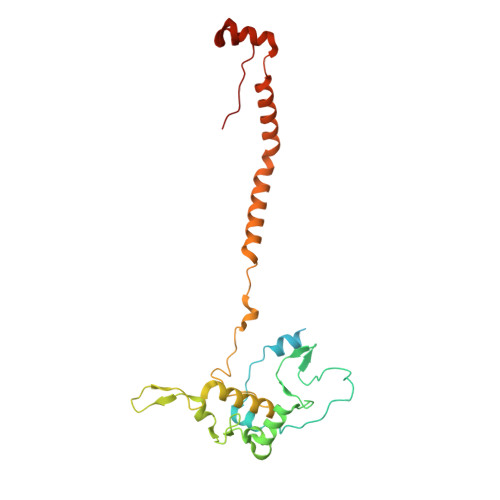
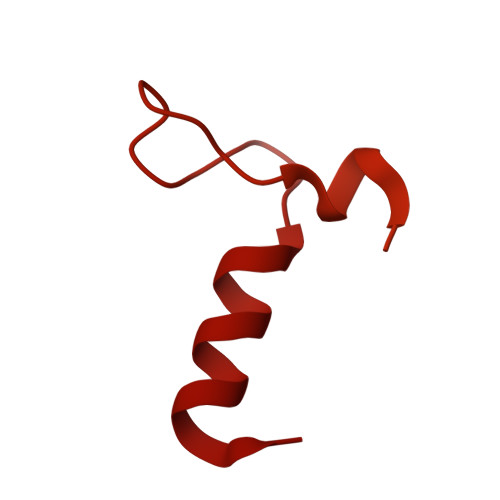
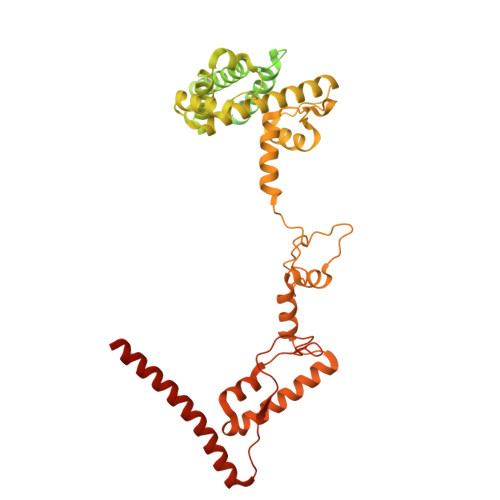
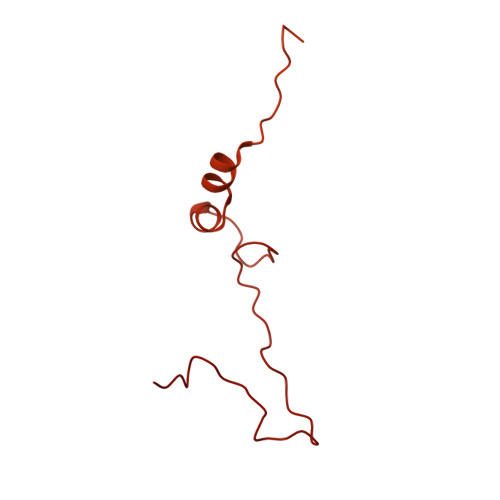
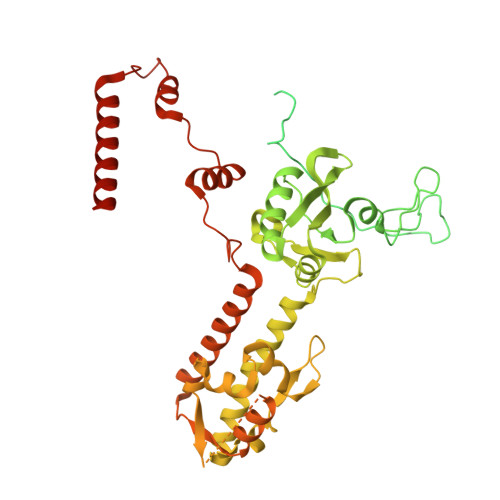
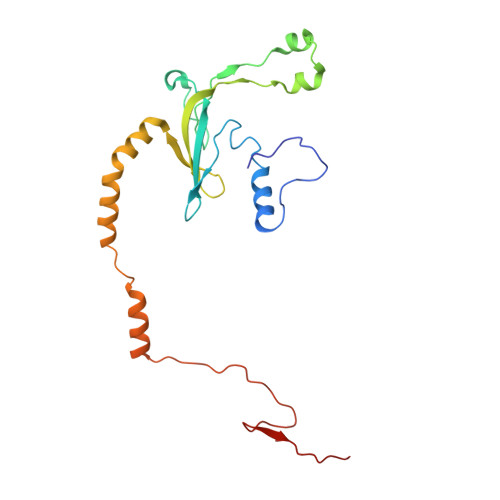
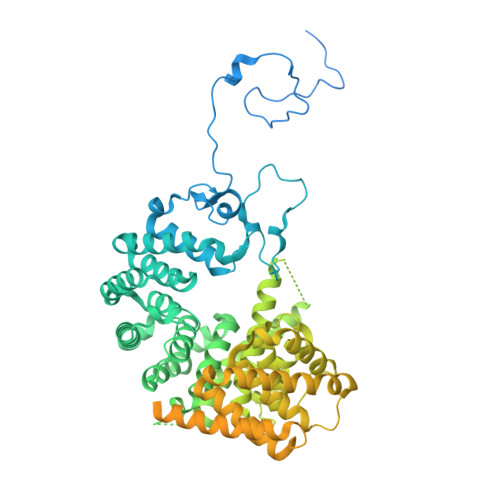
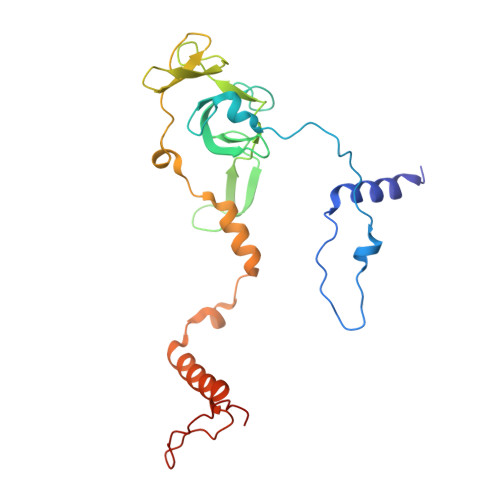
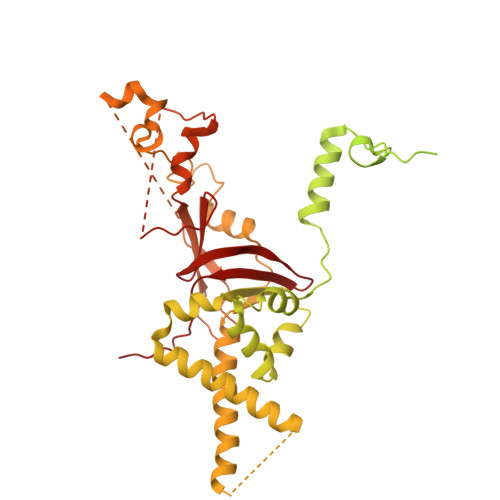
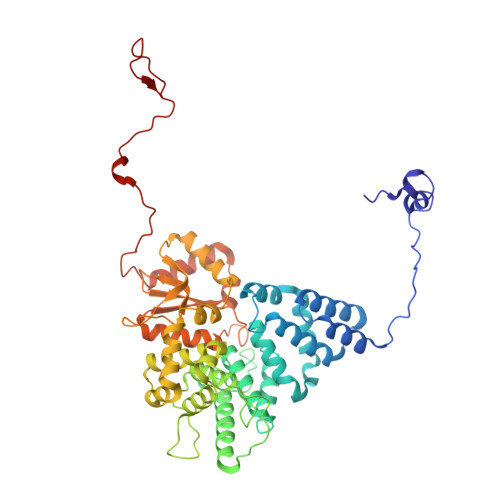
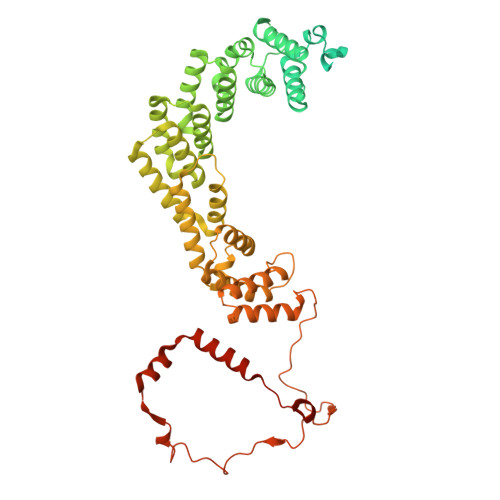
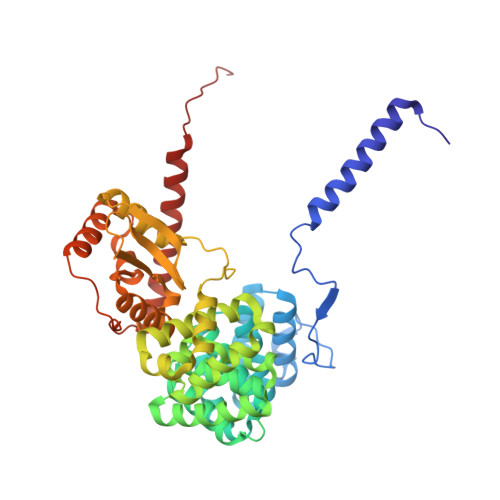
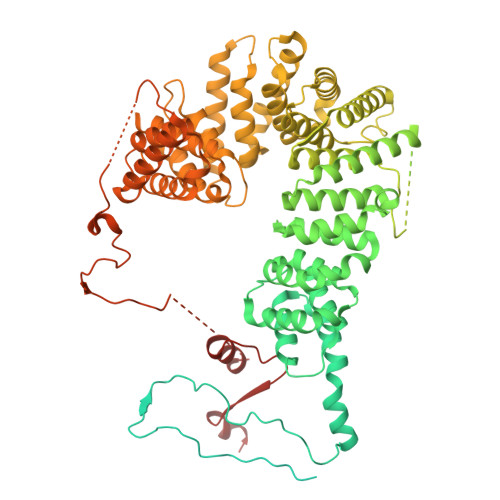
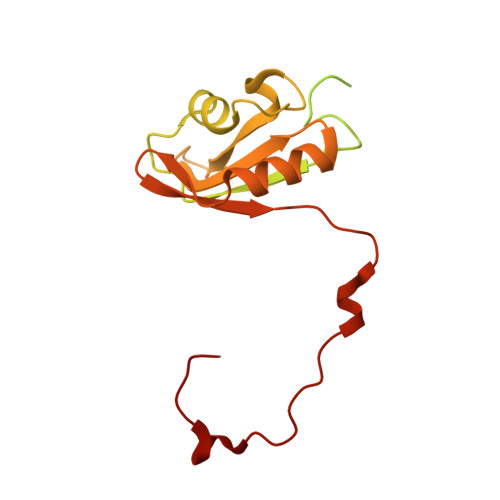
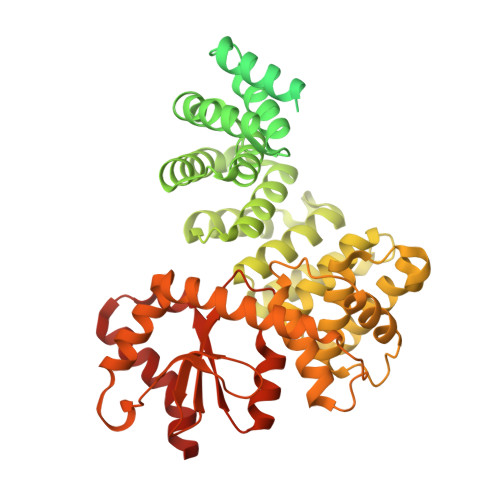
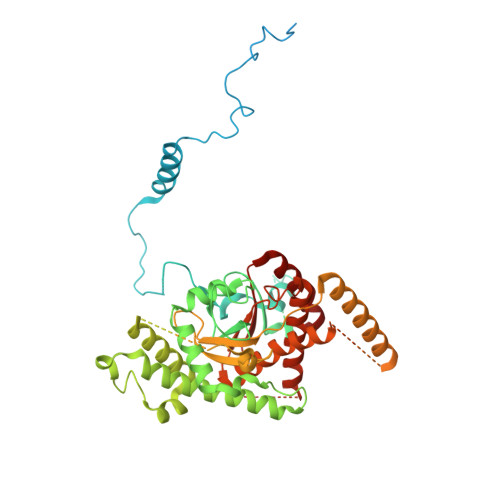
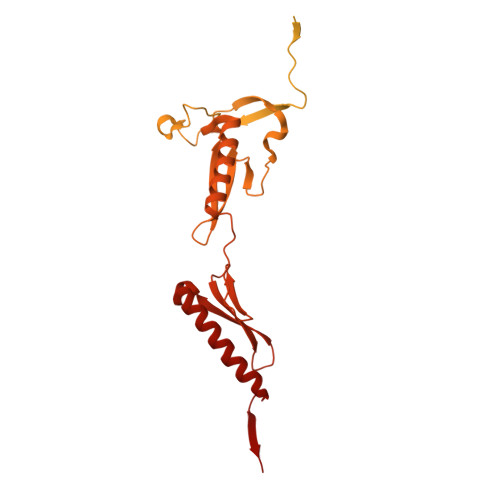
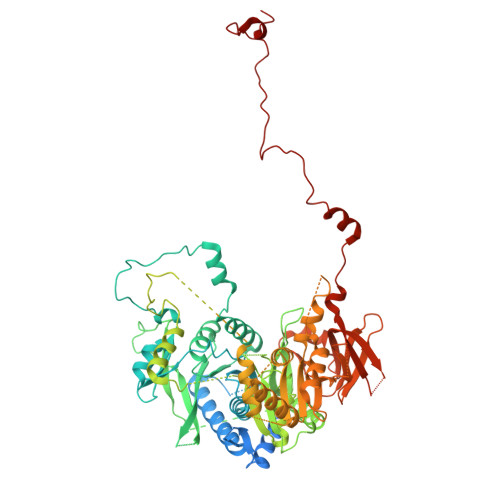
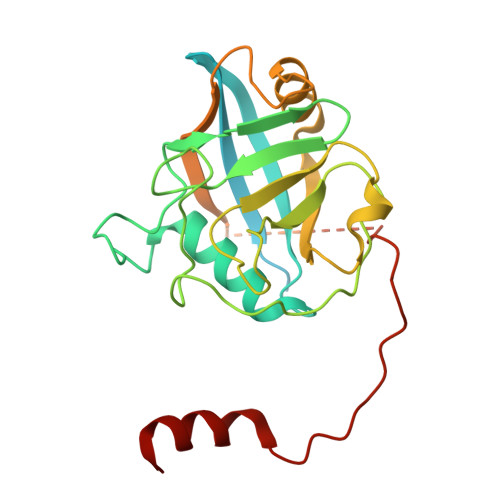
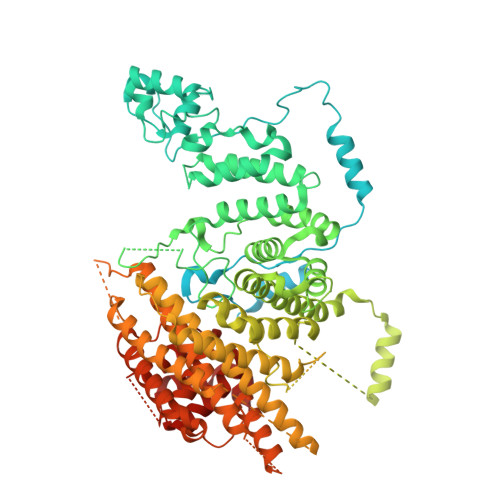
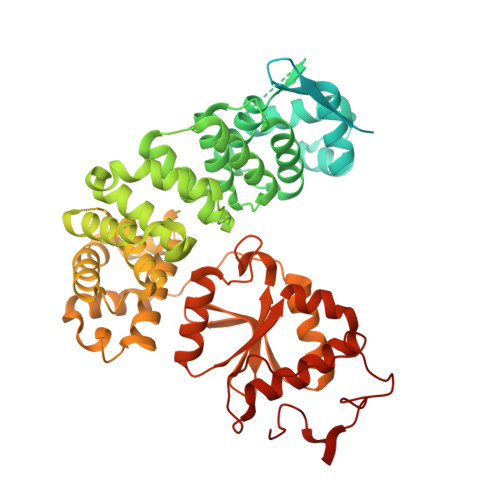
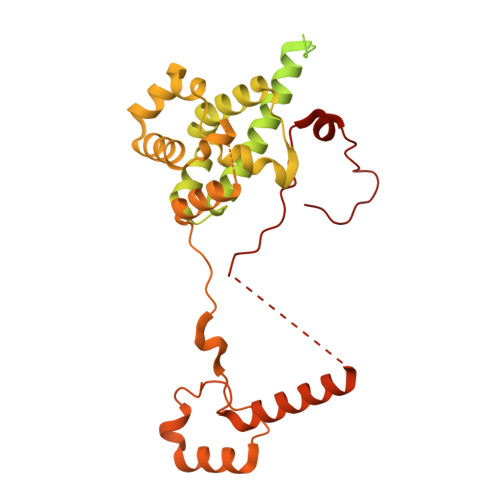
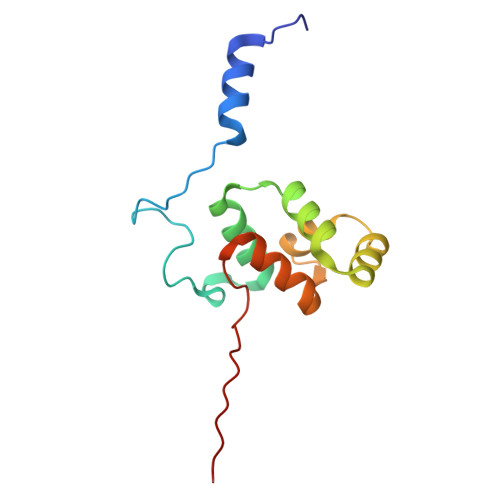
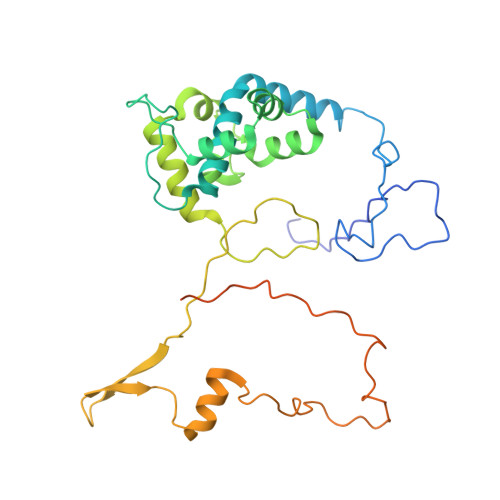
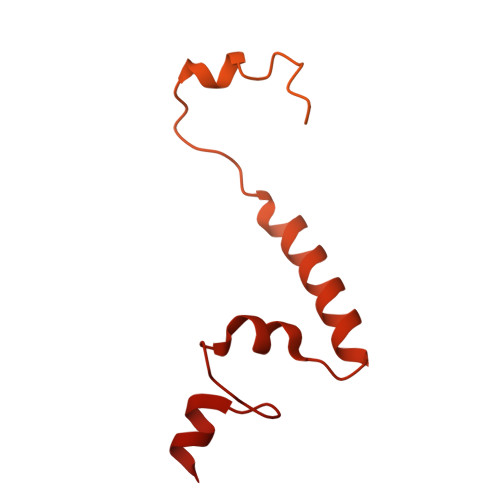
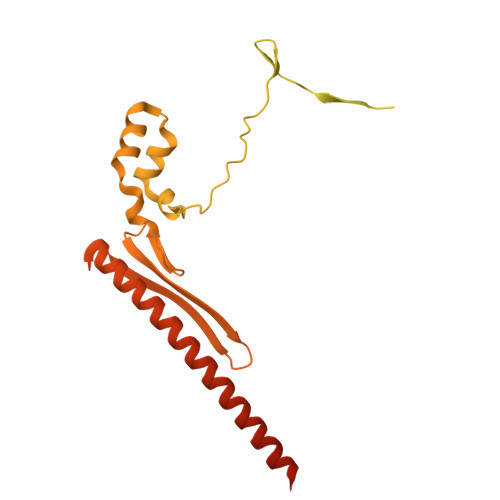
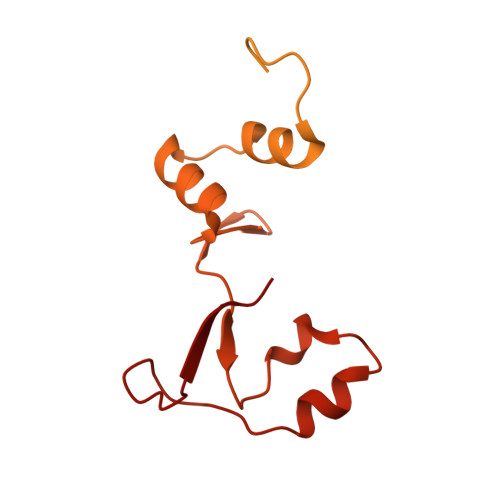
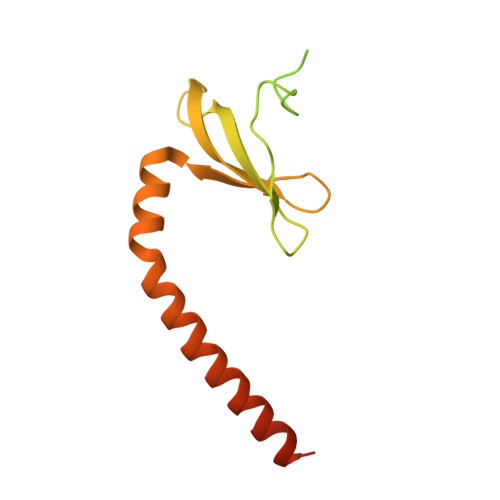
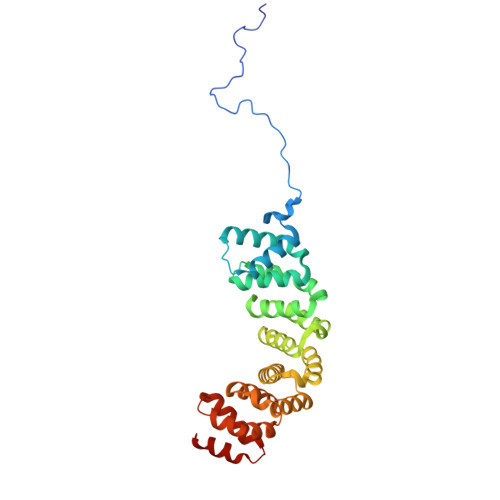
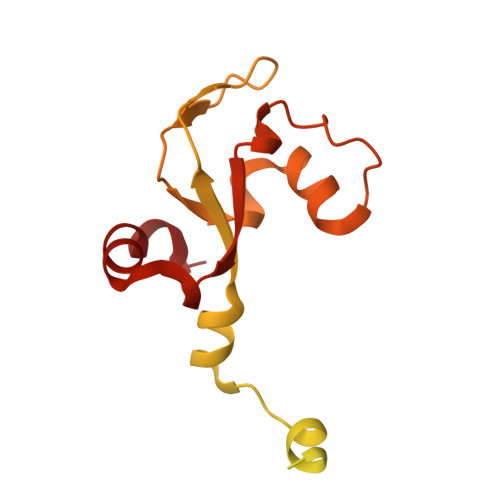
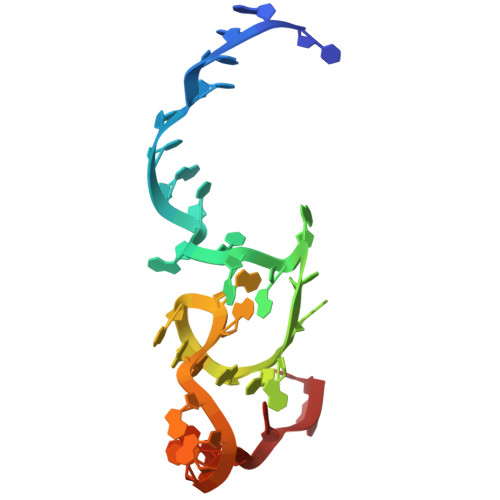
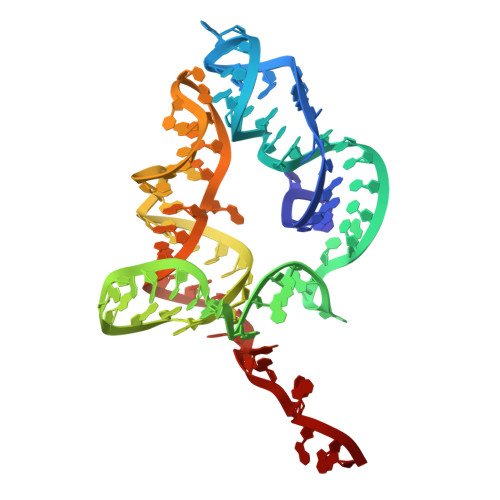
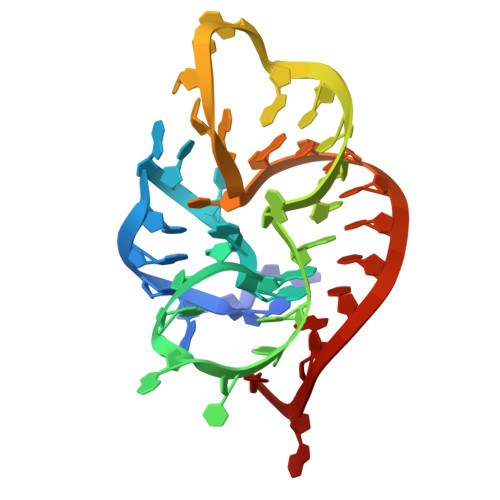
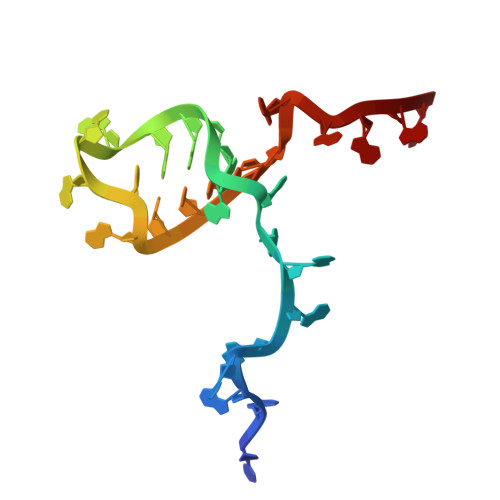
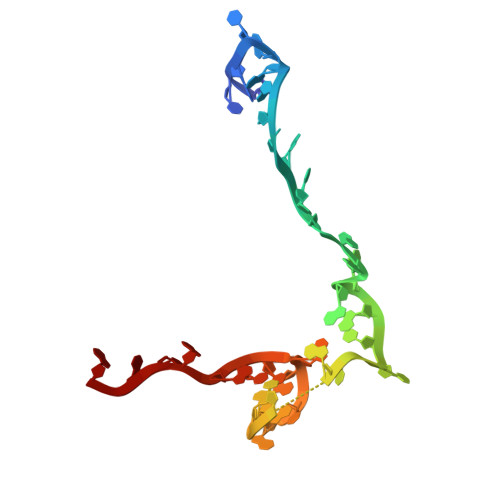
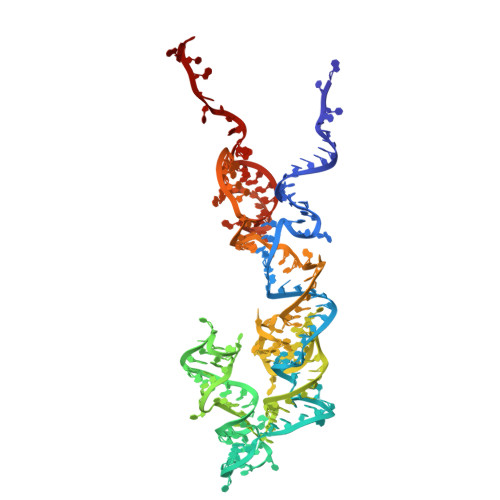
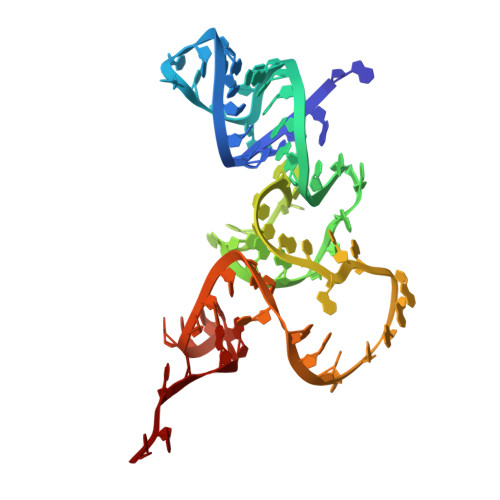
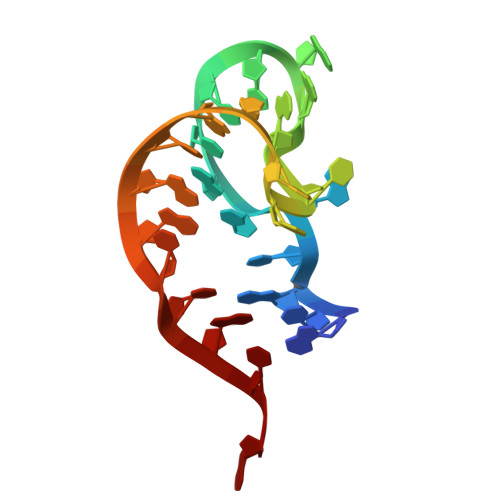
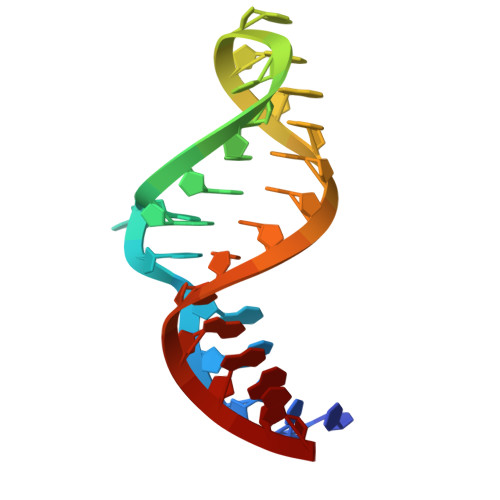
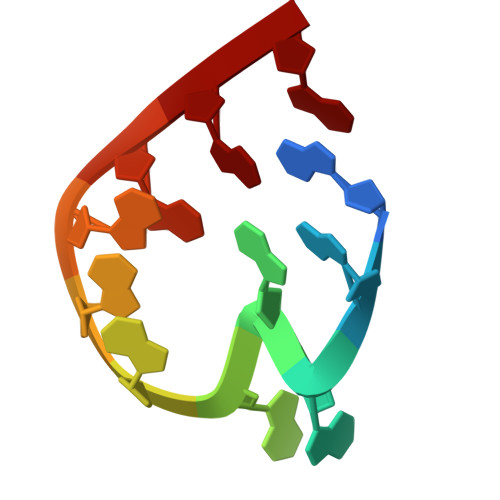
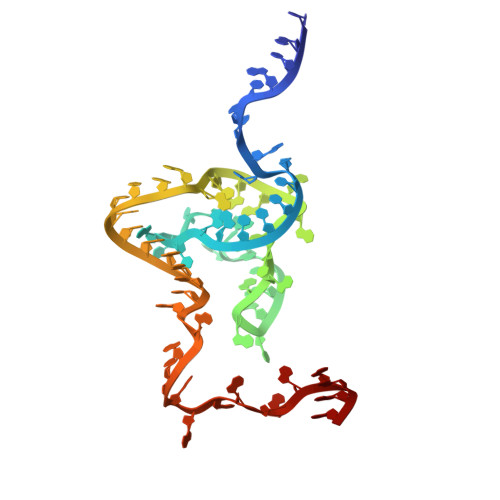
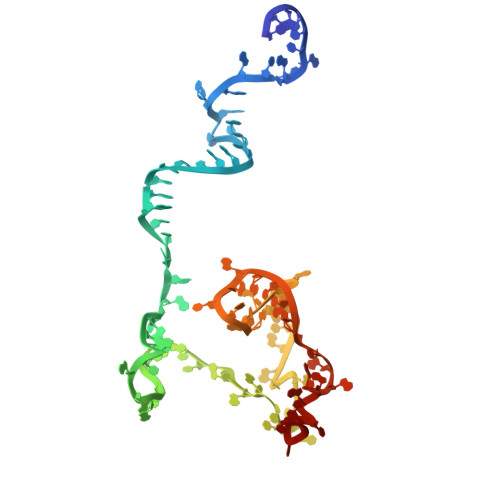
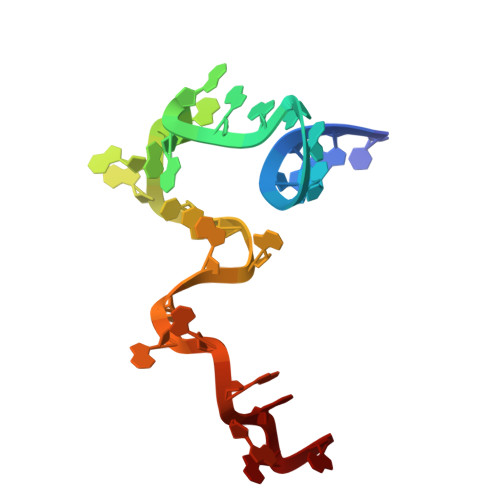
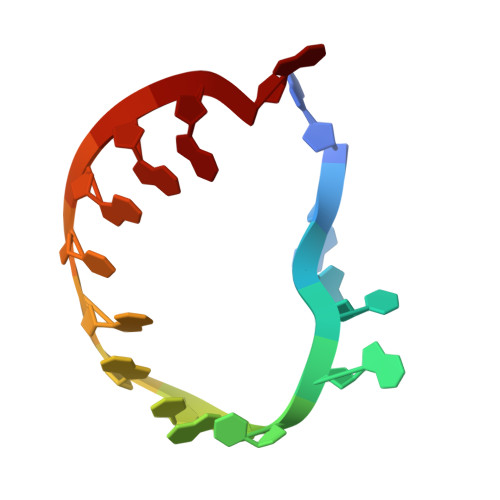
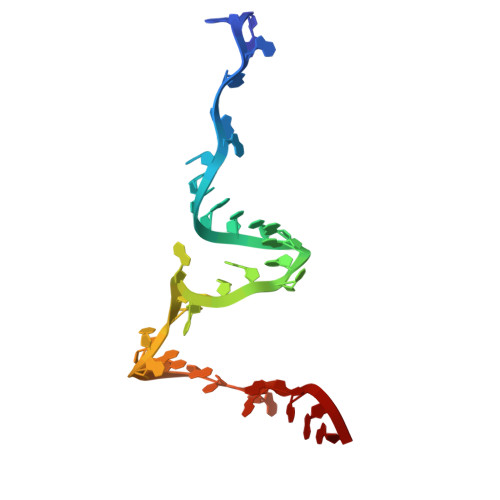
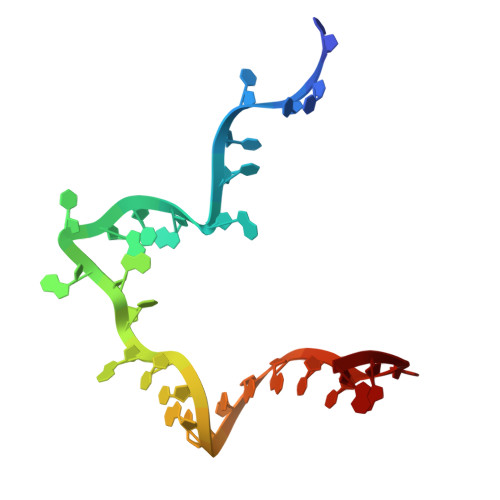
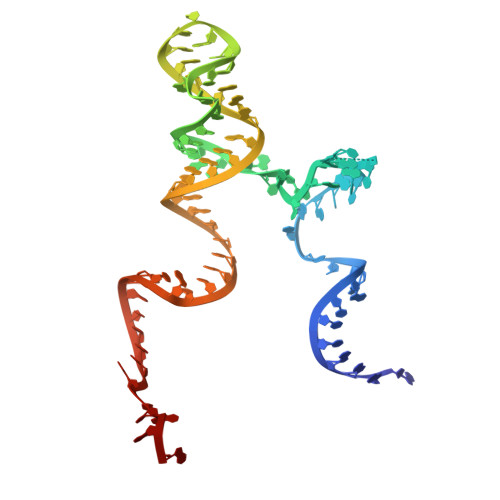
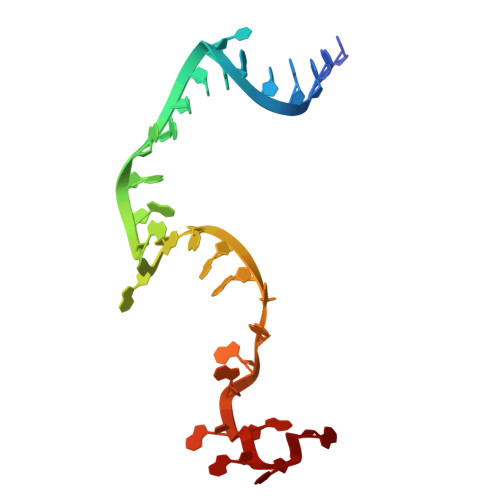
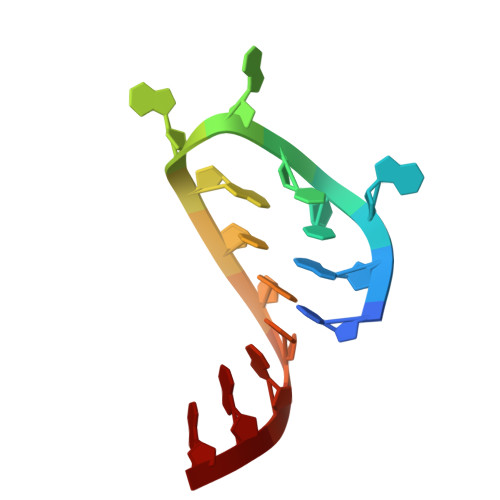
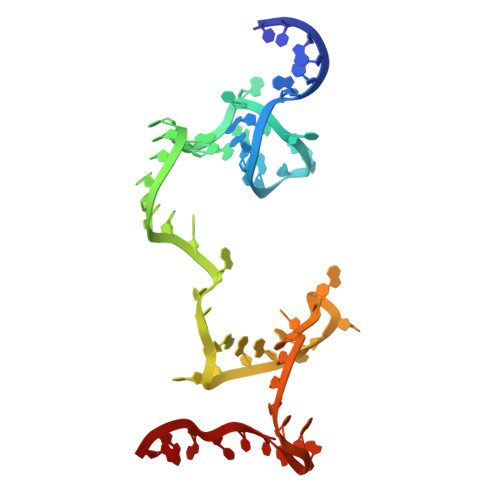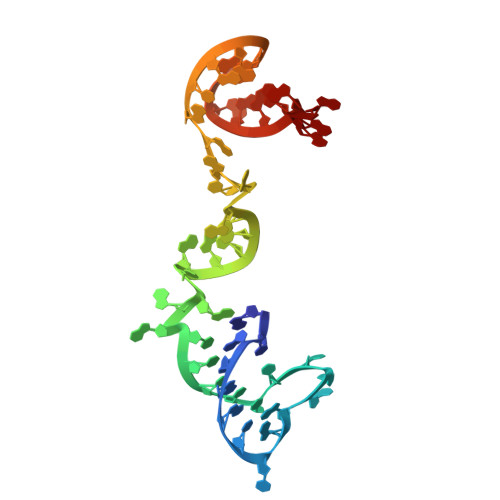Numerous rRNA molecules form the apicomplexan mitoribosome via repurposed protein and RNA elements.
Shikha, S., Tobiasson, V., Ferreira Silva, M., Ovciarikova, J., Beraldi, D., Muhleip, A., Sheiner, L.(2025) Nat Commun 16: 817-817
- PubMed: 39827269 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-56057-9
- Primary Citation Related Structures:
9HQV, 9I05 - PubMed Abstract:
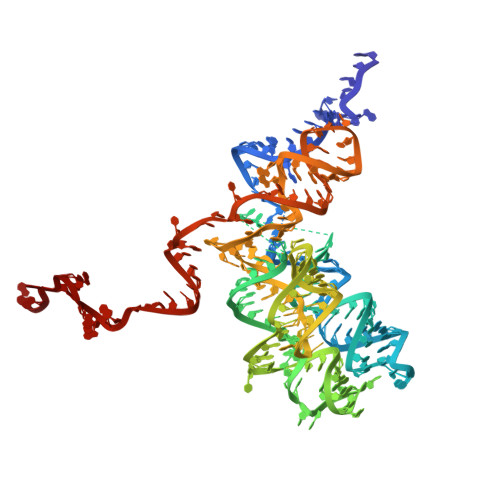
Mitochondrial ribosomes (mitoribosomes) are essential, and their function of synthesising mitochondrial proteins is universal. The core of almost all mitoribosomes is formed from a small number of long and self-folding rRNA molecules. In contrast, the mitoribosome of the apicomplexan parasite Toxoplasma gondii assembles from over 50 extremely short rRNA molecules. Here, we use cryo-EM to discover the features that enable this unusual mitoribosome to perform its function. We reveal that poly-A tails added to rRNA molecules are integrated into the ribosome, and we demonstrate their essentiality for mitoribosome formation and for parasite survival. This is a distinct function for poly-A tails, which are otherwise known primarily as stabilisers of messenger RNAs. Furthermore, while ribosomes typically consist of unique rRNA sequences, here nine sequences are used twice, each copy integrated in a different mitoribosome domain, revealing one of the mechanisms enabling the extreme mitochondrial genome reduction characteristic to Apicomplexa and to a large group of related microbial eukaryotes. Finally, several transcription factor-like proteins are repurposed to compensate for reduced or lost critical ribosomal domains, including members of the ApiAP2 family thus far considered to be DNA-binding transcription factors.
- School of Infection and Immunity, University of Glasgow, Glasgow, Scotland, UK.
Organizational Affiliation: