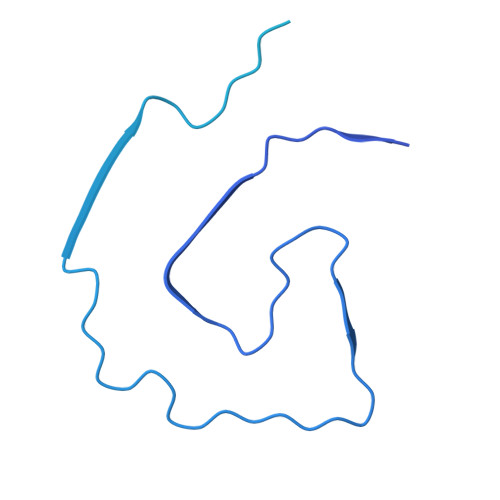
Apolipoprotein A-IV fibrils: structural diagnosis of mixed cardiac amyloidosis.
Aibara, S., Kassner, A., Wong, E., Klingel, K., Papworth, M., Althage, M., Wang, Q.D., Correia, C., Milting, H., de Oliveira, T.M.(2025) Nat Commun 16: 9276-9276
- PubMed: 41115976 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-64902-0
- Primary Citation Related Structures:
9HX3, 9HX4 - PubMed Abstract:
Cardiac amyloidosis (CA) occurs when misfolded proteins deposit as fibrils in the extracellular space of the heart. The fibrillogenic properties of apolipoprotein A-IV (ApoAIV) have been histologically observed and associated with CA pathogenesis. We report the structure of an ApoAIV amyloid from a patient's heart, which coexist amongst transthyretin (TTR) amyloids. These cases of undetected mixed CA highlight the importance of developing broad-spectrum anti-amyloid treatments to improve outcomes in patients.
- Protein Sciences, Structure, and Biophysics, Discovery Sciences, R&D, AstraZeneca, Cambridge, UK.
Organizational Affiliation: