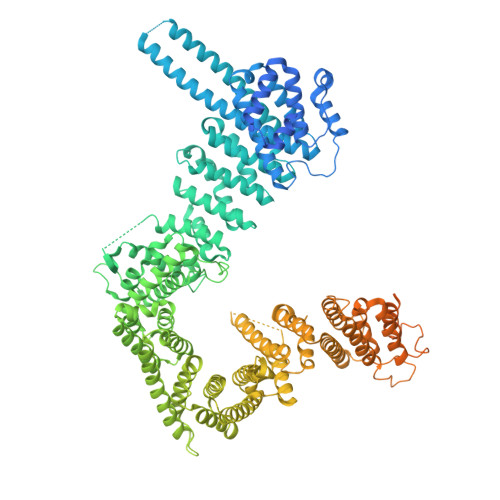
Substrate recognition by human separase.
Yu, J., Schmidt, S., Botto, M., Lee, K., Ghent, C.M., Goodfried, J.M., Howe, A., O'Reilly, F.J., Morgan, D.O., Boland, A.(2025) Sci Adv 11: eady9807-eady9807
- PubMed: 41223273 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.ady9807
- Primary Citation Related Structures:
9HM7, 9HMA, 9HMS, 9HMV, 9HN0, 9HN4, 9HN5 - PubMed Abstract:
The cohesin complex encircles sister chromatids in early mitosis. At anaphase onset, sister separation is triggered by the proteolytic cleavage of the cohesin subunit SCC1/RAD21 by separase. SCC1 contains two cleavage sites, where cleavage is stimulated by SCC1 phosphorylation. Substrate recognition and cleavage are only partly understood. Here, we determined structures of human separase in apo- or substrate-bound forms that, together with biochemical analysis, provide critical insights into separase cleavage regulation. We verify the first SCC1 cleavage site and reassign the second. We show that substrates, including separase autocleavage sites and the two SCC1 cleavage sites, interact with docking sites in separase, including five phosphate-binding sites. We also describe the interaction between the cohesin subunit SA1/SA2 and separase, which promotes cleavage at the second SCC1 site. Using cross-linking mass spectrometry and cryo-electron microscopy, we propose how cohesin is targeted by human separase. Our work provides an extensive functional and structural framework that explains a key event in cell division.
- Department of Molecular and Cellular Biology, University of Geneva, Geneva, Switzerland.
Organizational Affiliation: