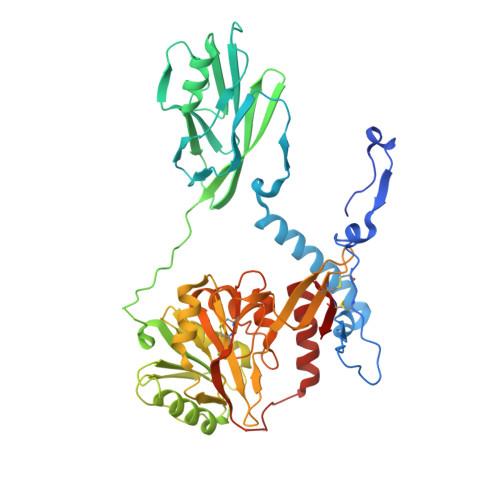
Conformational dynamics and membrane insertion mechanism of B4GALNT1 in ganglioside synthesis.
Welland, J.W.J., Barrow, H.G., Stansfeld, P.J., Deane, J.E.(2025) Nat Commun 16: 5442-5442
- PubMed: 40593514 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-60593-9
- Primary Citation Related Structures:
9H6J, 9H6K, 9H6L - PubMed Abstract:
Glycosphingolipids (GSLs) are crucial membrane components involved in essential cellular pathways. Complex GSLs, known as gangliosides, are synthesised by glycosyltransferase enzymes and imbalances in GSL metabolism cause severe neurological diseases. B4GALNT1 synthesises the precursors to the major brain gangliosides. Loss of B4GALNT1 function causes hereditary spastic paraplegia, while its overexpression is linked to cancers including childhood neuroblastoma. Here, we present crystal structures of the human homodimeric B4GALNT1 enzyme demonstrating dynamic remodelling of the substrate binding site during catalysis. We show that processing of lipid substrates by B4GALNT1 is severely compromised when surface loops flanking the active site are mutated from hydrophobic residues to polar. Molecular dynamics simulations support that these loops can insert into the lipid bilayer explaining how B4GALNT1 accesses and processes lipid substrates. By combining structure prediction and molecular simulations we propose that this mechanism of dynamic membrane insertion is exploited by other, structurally distinct GSL synthesising enzymes.
- Cambridge Institute for Medical Research, Department of Clinical Neuroscience, University of Cambridge, Cambridge, UK.
Organizational Affiliation: