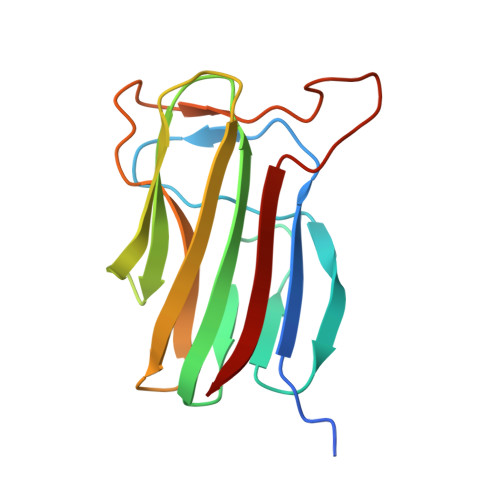
Cryo-EM analysis of the Bacillus thuringiensis extrasporal matrix identifies F-ENA as a widespread family of endospore appendages across the Firmicutes phylum.
Sleutel, M., Sogues, A., Gerven, N.V., Jonsmoen, U.L., Molle, I.V., Fislage, M., Theunissen, L.D., Bellis, N.F., Baquero, D.P., Egelman, E.H., Krupovic, M., Wang, F., Aspholm, M., Remaut, H.(2025) bioRxiv
- PubMed: 39990323 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2025.02.11.637640
- Primary Citation Related Structures:
9H38, 9H3D, 9I0Y - PubMed Abstract:
For over 100 years, Bacillus thuringiensis (Bt) has been used as an agricultural biopesticide to control pests caused by insect species in the orders of Lepidoptera, Diptera and Coleoptera. Under nutrient starvation, Bt cells differentiate into spores and associated toxin crystals that can adopt biofilm-like aggregates. We reveal that such Bt spore/toxin biofilms are embedded in a fibrous extrasporal matrix (ESM), and using cryoID, we resolved the structure and molecular identity of an uncharacterized type of pili, referred to here as Fibrillar ENdospore Appendages or 'F-ENA'. F-ENA are monomolecular protein polymers tethered to the exosporium of Bt and are decorated with a flexible tip fibrillum. Phylogenetic analysis reveals that F-ENA is widespread not only in the class Bacilli, but also in the class Clostridia, and the cryoEM structures of F-ENA filaments from Bacillus, Anaerovorax and Paenibaccilus reveal subunits with a generic head-neck domain structure, where the β-barrel neck of variable length latch onto a preceding head domain through short N-terminal hook peptides. In Bacillus , two collagen-like proteins (CLP) respectively tether F-ENA to the exosporium (F-Anchor), or constitute the tip fibrillum at the distal terminus of F-ENA (F-BclA). Sedimentation assays point towards F-ENA involvement in spore-spore clustering, likely mediated via F-BclA contacts and F-ENA bundling through the antiparallel interlocking of the head-neck units.
- Structural Biology Brussels, Vrije Universiteit Brussel, Pleinlaan 2, 1050 Brussels, Belgium.
Organizational Affiliation: