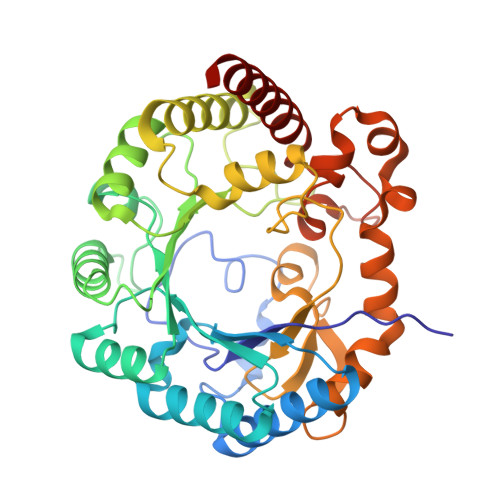
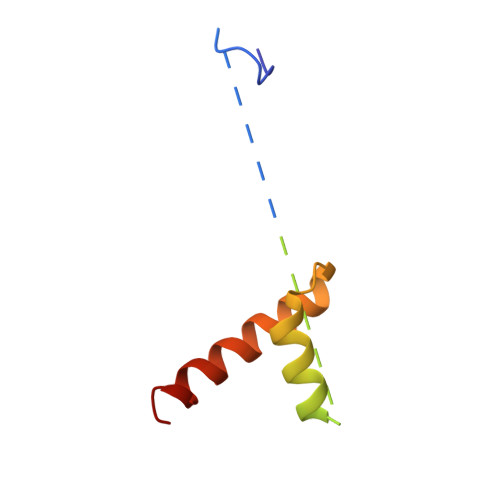
Peptide Recognition and Mechanism of the Radical S -Adenosyl-l-methionine Multiple Cyclophane Synthase ChlB.
Ruel, J., Nguyen, T.Q.N., Morishita, Y., Usclat, A., Martin, L., Amara, P., Kieffer-Jaquinod, S., Stefanoiu, M.C., de la Mora, E., Morinaka, B.I., Nicolet, Y.(2025) J Am Chem Soc 147: 16850-16863
- PubMed: 40354606 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.4c16004
- Primary Citation Related Structures:
9GM3, 9GMC - PubMed Abstract:
Ribosomally synthesized and post-translationally modified peptides (RiPPs) represent a valuable class of natural products, often featuring macrocyclization, which enhances stability and rigidity to achieve specific conformations, frequently underlying antibiotic activity. ChlB is a metalloenzyme with two catalytic domains─a radical S -adenosyl-l-methionine (SAM) domain and an α-ketoglutarate-dependent oxygenase─that work in tandem to sequentially form three cyclophanes and introduce three hydroxyl groups into its substrate peptide, ChlA. Here, we present the crystal structure of the radical SAM domain of ChlB in complex with ChlA, revealing the mechanism underlying cyclophane formation. These structures also elucidate how the leader sequence of ChlA interacts with ChlB. By combining structural, in vitro, and in vivo approaches, we determined the precise sequence of the three cyclophane formations, interspersed with hydroxylation events. Our findings demonstrate a back-and-forth movement of the core peptide between the radical SAM domain and the oxygenase domain, which drives the stepwise modification process, leading to the fully modified peptide.
- Univ. Grenoble Alpes, CEA, CNRS, IBS, Metalloproteins Unit, Grenoble F-38000, France.
Organizational Affiliation: