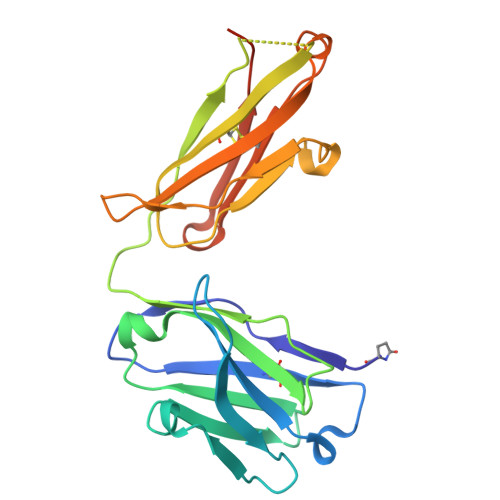
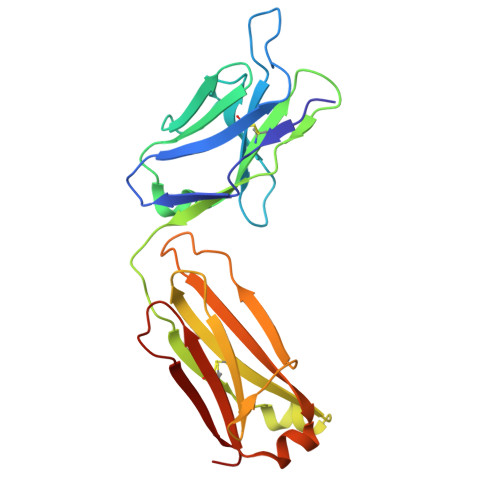
Improved targeted delivery of antisense oligonucleotide with an antibody mask.
Hsia, H.E., Zanini, C., Simonneau, C., Fraidling, J., Kraft, T.E., Mayer, K., Sommer, A., Indlekofer, A., Wirth, T., Benz, J., Georges, G., Langer, L.M., Gassner, C., Larraillet, V., Lohmann, S., Koller, E., Manso, M., Ravn, J., Hofer, K., Emrich, T., Niewohner, J., Schumacher, F., Brinkmann, U.(2025) Nucleic Acids Res 53
- PubMed: 41166443 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaf487
- Primary Citation Related Structures:
9GF5, 9GFD, 9GFJ, 9GFL - PubMed Abstract:
Antisense oligonucleotides (ASOs) are synthetic nucleic acid strands designed to modulate gene expression by binding to RNA transcripts. In brain diseases, transferrin receptor (TfR)-targeting antibody-ASO conjugates have shown promise for brain delivery of ASOs via transcytosis in preclinical studies. This enables the more patient friendly intravenous or subcutaneous administration of the compounds. However, these conjugates can exhibit faster plasma clearance and different peripheral pharmacokinetic profiles due to ASO modifications, such as phosphorothioate (PS) linkages and locked nucleic acids (LNAs). In this study, we employed an antibody recognizing LNA- and PS-modified ASOs independent of the base sequences as a cloaking module to mitigate these issues. Using Kutzneria albida transglutaminase (KTG) technology and click chemistry, we generated TfR antibody-ASO conjugates with covalently or noncovalently incorporated ASO binders. Additionally, a noncovalent carrier antibody approach was explored. These conjugates and complexes with additional ASO binder(s) show improved TfR-targeted cellular uptake, undergo transcytosis in cellular blood-brain barrier models, show less nonspecific cellular accumulation, and anticipated ASO activity.
- Roche Pharma Research and Early Development (pRED), Roche Innovation Center Munich, Roche Diagnostics GmbH, Penzberg82377, Germany.
Organizational Affiliation: