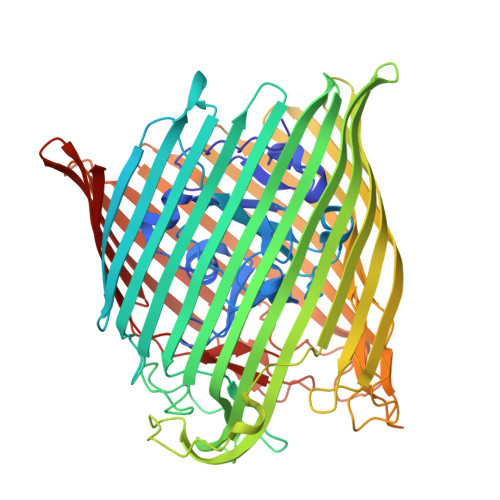
Structure of the Outer Membrane Transporter FemA and Its Role in the Uptake of Ferric Dihydro-Aeruginoic Acid and Ferric Aeruginoic Acid in Pseudomonas aeruginosa .
Will, V., Moynie, L., Si Ahmed Charrier, E., Le Bas, A., Kuhn, L., Volck, F., Chicher, J., Aksoy, H., Madec, M., Antheaume, C., Mislin, G.L.A., Schalk, I.J.(2025) ACS Chem Biol 20: 690-706
- PubMed: 40035455 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.4c00820
- Primary Citation Related Structures:
8S34, 9EX3, 9F2T, 9FVQ - PubMed Abstract:
Iron is essential for bacterial growth, and Pseudomonas aeruginosa synthesizes the siderophores pyochelin (PCH) and pyoverdine to acquire it. PCH contains a thiazolidine ring that aids in iron chelation but is prone to hydrolysis, leading to the formation of 2-(2-hydroxylphenyl)-thiazole-4-carbaldehyde (IQS). Using mass spectrometry, we demonstrated that PCH undergoes hydrolysis and oxidation in solution, resulting in the formation of aeruginoic acid (AA). This study used proteomic analyses and fluorescent reporters to show that AA, dihydroaeruginoic acid (DHA), and PCH induce the expression of femA , a gene encoding the ferri-mycobactin outer membrane transporter in P. aeruginosa . Notably, the induction by AA and DHA was observed only in strains unable to produce pyoverdine, suggesting their weaker iron-chelating ability compared to that of pyoverdine. 55 Fe uptake assays demonstrated that both AA-Fe and DHA-Fe complexes are transported via FemA; however, no uptake was observed for PCH-Fe through this transporter. Structural studies revealed that FemA is able to bind AA 2 -Fe or DHA 2 -Fe complexes. Key interactions are conserved between FemA and these two complexes, with specificity primarily driven by one of the two siderophore molecules. Interestingly, although no iron uptake was noted for PCH through FemA, the transporter also binds PCH-Fe in a similar manner. These findings show that under moderate iron deficiency, when only PCH is produced by P. aeruginosa , degradation products AA and DHA enhance iron uptake by inducing femA expression and facilitating iron transport through FemA. This provides new insights into the pathogen's strategies for iron homeostasis.
- CNRS, University of Strasbourg, UMR7242, UMR7242, ESBS, Bld Sébastien Brant, F-67412 Strasbourg, Illkirch, France.
Organizational Affiliation: