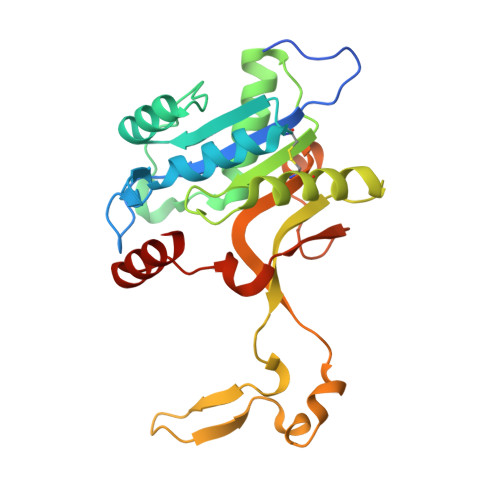
Structural dissection of the CMP-pseudaminic acid synthetase, PseF.
Keenan, T., Cowan, A.R., Flack, E.K.P., Hatton, N.E., Walklett, A.J., Thomas, G.H., Hemsworth, G.R., Fascione, M.A.(2024) Structure 32: 2399
- PubMed: 39393361 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2024.09.017
- Primary Citation Related Structures:
9FTB, 9FTC - PubMed Abstract:
Pseudaminic acid is a non-mammalian sugar found in the surface glycoconjugates of many bacteria, including several human pathogens, and is a virulence factor thought to facilitate immune evasion. The final step in the biosynthesis of the nucleotide activated form of the sugar, CMP-Pse5Ac7Ac is performed by a CMP-Pse5Ac7Ac synthetase (PseF). Here we present the biochemical and structural characterization of PseF from Aeromonas caviae (AcPseF), with AcPseF displaying metal-dependent activity over a broad pH and temperature range. Upon binding to CMP-Pse5Ac7Ac, AcPseF undergoes dynamic movements akin to other CMP-ulosonic acid synthetases. The enzyme clearly discriminates Pse5Ac7Ac from other ulosonic acids, through active site interactions with side-chain functional groups and by positioning the molecule in a hydrophobic pocket. Finally, we show that AcPseF binds the CMP-Pse5Ac7Ac side chain in the lowest energy conformation, a trend that we observed in the structures of other enzymes of this class.
- Department of Chemistry, University of York, York YO10 5DD, UK.
Organizational Affiliation: