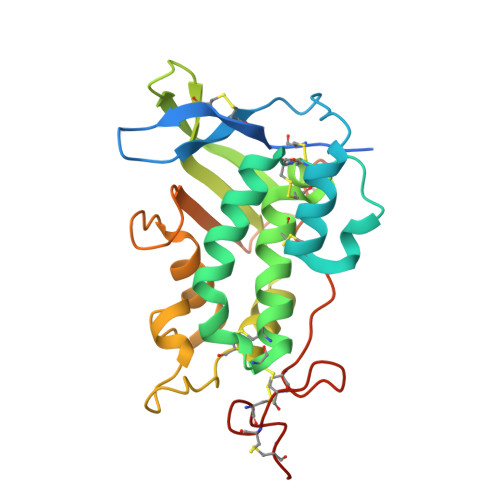
Structural and biochemical analysis of ligand binding in yeast Niemann-Pick type C1-related protein.
Nel, L., Thaysen, K., Jamecna, D., Olesen, E., Szomek, M., Langer, J., Frain, K.M., Hoglinger, D., Wustner, D., Pedersen, B.P.(2025) Life Sci Alliance 8
- PubMed: 39455279 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.202402990
- Primary Citation Related Structures:
9F40, 9F41 - PubMed Abstract:
In eukaryotes, integration of sterols into the vacuolar/lysosomal membrane is critically dependent on the Niemann-Pick type C (NPC) system. The system consists of an integral membrane protein, called NCR1 in yeast, and NPC2, a luminal soluble protein that transfers sterols to the N-terminal domain (NTD) of NCR1 before membrane integration. Both proteins have been implicated in sterol homeostasis of yeast and humans. Here, we investigate sterol and lipid binding of the NCR1/NPC2 transport system and determine crystal structures of the sterol binding NTD. The NTD binds both ergosterol and cholesterol, with nearly identical conformations of the binding pocket. Apart from sterols, the NTD can also bind fluorescent analogs of phosphatidylinositol, phosphatidylcholine, and phosphatidylserine, as well as sphingosine and ceramide. We confirm the multi-lipid scope of the NCR1/NPC2 system using photo-crosslinkable and clickable lipid analogs, namely, pac-cholesterol, pac-sphingosine, and pac-ceramide. Finally, we reconstitute the transfer of pac-sphingosine from NPC2 to the NTD in vitro. Collectively, our results support that the yeast NPC system can work as versatile machinery for vacuolar homeostasis of structurally diverse lipids, besides ergosterol.
- https://ror.org/01aj84f44 Department of Molecular Biology and Genetics, Aarhus University, Aarhus, Denmark.
Organizational Affiliation: