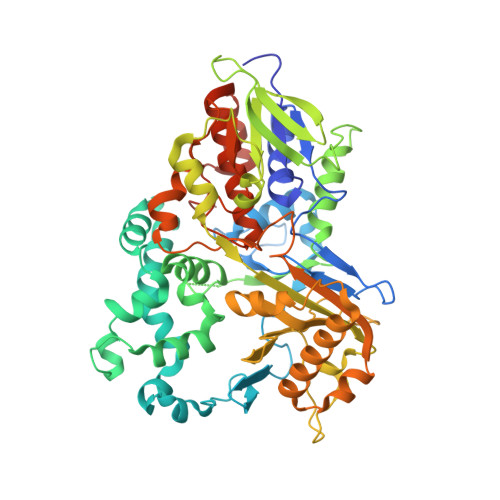
Crystal structure and enzyme engineering of the broad substrate spectrum l-amino acid oxidase 4 from the fungus Hebeloma cylindrosporum.
Koopmeiners, S., Gilzer, D., Widmann, C., Berelsmann, N., Spross, J., Niemann, H.H., Fischer von Mollard, G.(2024) FEBS Lett 598: 2306-2320
- PubMed: 39152524 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.15002
- Primary Citation Related Structures:
9ENH, 9ENI, 9ENJ, 9ENK, 9ENN - PubMed Abstract:
l-Amino acid oxidases (LAAOs) catalyze the oxidative deamination of l-amino acids to α-keto acids. Recombinant production of LAAOs with broad substrate spectrum remains a formidable challenge. We previously achieved this for the highly active and thermostable LAAO4 of Hebeloma cylindrosporum (HcLAAO4). Here, we crystallized a proteolytically truncated surface entropy reduction variant of HcLAAO4 and solved its structure in substrate-free form and in complex with diverse substrates. The ability to support the aliphatic portion of a substrate's side chain by an overall hydrophobic active site is responsible for the broad substrate spectrum of HcLAAO4, including l-amino acids with big aromatic, acidic and basic side chains. Based on the structural findings, we generated an E288H variant with increased activity toward pharmaceutical building blocks of high interest.
- Biochemistry III, Department of Chemistry, Bielefeld University, Bielefeld, Germany.
Organizational Affiliation: